Introduction:
A Semaglutide Sameness Study is a comprehensive analytical comparison between a proposed generic semaglutide and the reference product. The goal is to prove they are identical in sequence, composition, and structure. For semaglutide (a 31-amino-acid GLP-1 analog), even minor sequence variants or new impurities can affect efficacy or safety. Therefore, regulators (FDA, Health Canada, etc.) require robust orthogonal evidence of sameness beyond just bioequivalence. Our experts at ResolveMass Laboratories use state-of-the-art techniques to compare the generic and RLD side-by-side, covering:
- Peptide sequence confirmation: Verifying the exact amino acid sequence, including any modifications or conjugations.
- Molecular weight verification: Measuring the intact peptide mass to ensure it matches the theoretical mass.
- Impurity and degradation profiling: Identifying process- or storage-related impurities (truncations, oxidations, etc.) in each sample.
- Structural integrity (higher-order structure): Using NMR and other methods to confirm correct folding and disulfide bridges.
- Comparative analytical characterization: Side-by-side data overlays and statistical comparisons with the RLD.
By combining these analyses, we provide scientifically defensible proof of sameness. This evidence is essential for generic ANDA approval and ensures patient safety and regulatory compliance. Our multidisciplinary team’s deep peptide chemistry expertise and validated protocols deliver high-confidence results.
Summary:
- A Semaglutide Sameness Study rigorously demonstrates that a generic semaglutide is structurally and chemically identical to the reference listed drug (RLD).
- Key analyses include peptide sequencing and mapping via LC-MS/MS to confirm the exact amino acid order.
- Impurity profiling using high-resolution LC-MS/HRMS and HPLC detects any process- or degradation-related impurities, ensuring equivalence to the RLD.
- 1D/2D NMR spectroscopy provides orthogonal confirmation of semaglutide’s higher-order structure and disulfide bonds.
- ResolveMass Laboratories’ experienced team, advanced instruments, and validated workflows ensure regulatory-grade data and support a strong ANDA submission.
1. Peptide Sequencing by HRMS/MS
1.1 Objective
The goal of peptide sequencing is to confirm the complete primary structure of semaglutide, including the Aib substitution and the lipidation on Lys26. In practical terms, we must generate MS/MS fragmentation that covers the entire 31-amino-acid sequence and decisively identifies the modified residues. In a sameness study, the MS/MS spectra from the generic must align 100% with those from the RLD.
1.2 Structural Considerations
Several features of semaglutide affect the MS/MS approach. The non-natural amino acid Aib (α-aminobutyric acid) differs from alanine by an additional methyl group; it has the same nominal mass as valine, but in practice we treat Aib as a distinct residue in our sequence database. The bulky C18 diacid on Lys26 adds a large hydrophobic tail (~266 Da for stearic diacid) and may reduce peptide solubility or suppress fragmentation at that site. There are no disulfide bonds in semaglutide, but its strong helical tendencies and hydrophobic groups can alter ionization efficiencies. We account for these by using optimized solvent systems (e.g. HFIP or DMSO if needed) and by choosing proteases that maximize sequence coverage around modified sites.
1.3 Enzymatic Digestion Strategy
A single protease is unlikely to cover semaglutide fully due to the Lys26 acylation. Trypsin would normally cleave at Lys or Arg, but Lys26 is blocked by the lipid, so tryptic digestion yields large fragments. Therefore, we use a multi-enzyme approach:
- Glu-C: Cleaves C-terminal to Glu/Asp residues; this enzyme is not hindered by the Lys26 modification and can produce overlapping peptides across most of the chain.
- Chymotrypsin: Targets aromatic and hydrophobic residues, useful for generating fragments in regions with many nonpolar residues.
- Asp-N (optional): Cleaves N-terminal to Asp; provides an orthogonal cleavage pattern.
By combining these enzymes (often in parallel digests), we obtain overlapping peptide fragments that cover every region of semaglutide, including the N-terminus, the Aib site, and the lipidated Lys.
1.4 Experimental Procedure
A typical sequencing workflow includes the following steps:
- Buffer exchange: Semaglutide samples (RLD and generic) are buffer-exchanged into an MS-compatible digestion buffer (e.g. 50 mM ammonium bicarbonate).
- Reduction/alkylation: Although semaglutide has no disulfide bonds, a reduction step (DTT) and optional alkylation (iodoacetamide) are sometimes performed for consistency with other peptide protocols.
- Enzymatic digestion: Add protease at a 1:20–1:50 enzyme:substrate ratio and incubate at 37 °C for 4–16 hours. Each digestion (e.g. Glu-C, chymotrypsin) is performed separately.
- Quenching: Stop the reaction by lowering pH (e.g. adding formic acid to 0.1–1%). This also helps in activating the MS.
- LC-MS/MS analysis: Inject digests on an LC-HRMS/MS system (e.g. QTOF or Orbitrap) with a gradient optimized for peptide separation. Collect high-resolution full MS and MS/MS (data-dependent or targeted) for the expected masses.
1.5 Data Analysis
Data analysis is performed by searching the MS/MS spectra against a custom semaglutide sequence. The database includes the exact 31-residue sequence with variable modifications defined for Aib and the Lys26 lipid conjugate. For example, Lys26 can be defined with a +C18 diacid mass shift. We look for b- and y-ions that span the entire sequence, paying special attention to fragments containing Aib or the modified Lysine. Key metrics are:
- Sequence coverage: We aim for >95% coverage of the peptide chain from the combined enzyme digests.
- Mass accuracy: All fragment ions should match theoretical masses within tight tolerance (typically <5 ppm for HRMS).
- Identification of modifications: Peptides containing Aib or acylated Lys26 must be confidently identified by their unique mass shifts.
- Absence of unexpected variants: No additional peaks (indicative of impurities or sequence errors) should appear in the spectra.
Quantitatively, our acceptance criteria include exact mass matching (<5 ppm) for all fragments and identification of all predicted modified peptides. Even a single amino-acid discrepancy is unacceptable under FDA peptide sameness standards.
1.6 Key Observations / Acceptance Criteria
In the semaglutide sequencing data, we expect the following:
- Full sequence confirmation: All portions of the 31-residue sequence must be accounted for by identified peptides.
- Exact mass matching: The intact mass of semaglutide (including Aib and C18 acylation) matches the theoretical mass within the HRMS tolerance. Fragment ions for each peptide also match within tolerance.
- Modified peptide identification: The peptide containing Lys26 should show the extra mass of the lipid chain. All MS/MS ions that include the modified Lysine should align with calculated values.
- No unexpected variants: There should be no MS/MS peaks corresponding to any sequence variation or extra modification. A 100% alignment with the RLD’s fragmentation pattern is required.
If all criteria are met (complete sequence coverage, correct modification assignments, and no foreign peptides), we conclude that the primary structure of the generic semaglutide is identical to the RLD.
2. Peptide Mapping (Comparative Fingerprinting)
2.1 Objective
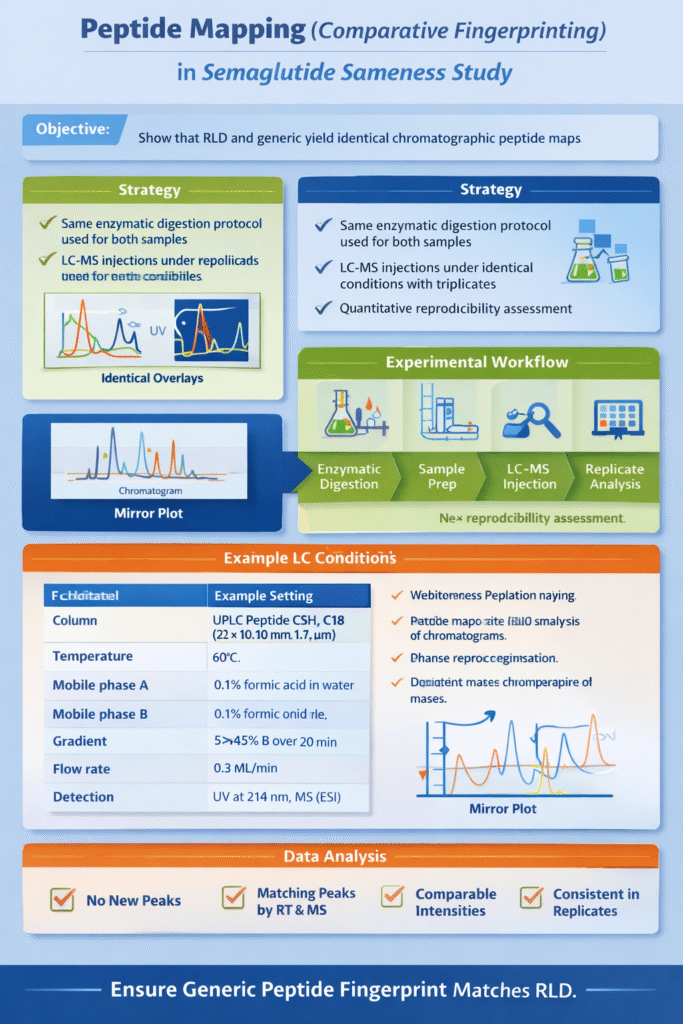
Peptide mapping provides a visual fingerprint to compare the generic and RLD semaglutide. The goal is to demonstrate that both samples yield identical chromatographic peptide maps when digested under the same conditions. If the generic is truly the same, the overlays of UV or MS chromatograms from RLD and test should show matching peaks (in retention time and MS identity) with no new or missing signals.
2.2 Strategy
To ensure reproducibility, we use the same digestion protocol on both the RLD and generic samples (same enzyme(s), buffer, time, and temperature). This creates a comparable set of peptides. The workflow is:
- Digest both samples (often in parallel batches).
- Perform LC-MS under identical conditions (same column, gradient, temperature, injection volume, etc.).
- Include at least triplicate injections of each to assess variability.
The digests are typically shorter (e.g. 4–6 hours) than sequencing digests to produce a consistent peptide profile. By aligning replicate runs, we confirm method consistency.
2.3 Experimental Workflow
- Enzymatic digestion: Using the chosen enzyme (or enzyme mix), digest RLD and generic side-by-side.
- Sample preparation: After quenching, dilute or buffer-exchange into the starting mobile phase if needed.
- LC-MS injection: Inject digests on an LC system equipped with a reversed-phase C18 column, using the same gradient for both samples.
- Detection: Monitor the eluent by UV at 214 nm (peptide bonds absorb strongly here) and by MS full-scan/fragmentation.
Replicate injections (e.g. n=3) are analyzed to ensure each peptide peak is consistently observed. Software can generate mirror plots of chromatograms to directly compare RLD vs generic (see below).
2.4 LC Conditions
A typical LC setup for a hydrophobic peptide like semaglutide uses a peptide-optimized C18 column (e.g. 2.1 mm × 100 mm, 1.7 µm particle size) at elevated temperature (~60–65 °C) to sharpen peaks. Mobile phases are 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile (B). We run a gradient (e.g. 5% to 50% B over 10–30 min) that is long enough to separate the lipidated peptides. Detection is dual: a tunable UV detector at 214 nm for peptide bonds and the MS for mass confirmation. For example, Waters showed UV detection at 214 nm successfully resolving peptide impurities.
Table 1. Example LC conditions for peptide mapping.
Parameter Example Setting Column UPLC Peptide CSH C18 (2.1×100 mm, 1.7 µm) Temperature 60 °C Mobile phase A 0.1% formic acid in water Mobile phase B 0.1% formic acid in acetonitrile Gradient 5→45% B over 20 min Flow rate 0.3 mL/min Detection UV at 214 nm, MS (ESI)
2.5 Data Analysis
After acquisition, we align chromatograms of RLD vs generic. Software (e.g. Waters UNIFI or Thermo Xcalibur) can overlay UV or MS traces. Key steps are:
- Peak alignment: Match peaks by retention time (RT) and MS mass.
- Overlay analysis: Create mirror plots of UV chromatograms; identical patterns indicate the same peptides. For instance, Waters demonstrated that a mirror plot of the sample and reference peptide maps showed a one-to-one correspondence of peaks.
- Quantitative comparison: Compare the area of each corresponding peak. Minor differences (±10–20%) in intensity are often acceptable, but major new peaks must be investigated.
2.6 Key Observations / Acceptance Criteria
In the peptide maps for semaglutide, the acceptance criteria are:
- Matching peaks: Every major peak in the RLD chromatogram must appear in the generic at the same RT (within experimental error, e.g. <0.1 min shift). Each should have the same MS identity (same peptide sequence).
- No new peaks: The generic should not show any additional peptide peaks above background. (Low-abundance differences are evaluated case-by-case.)
- Comparable intensities: The relative intensity pattern of peaks (area % distribution) should be similar. Minor variations due to injection or detector noise are tolerable.
- Reproducibility: Replicates of each sample must show consistent profiles.
Passing these criteria means the generic’s peptide fingerprint is indistinguishable from the RLD. In practice, if a mirror-plot overlay shows a perfect match of UV peaks (as in), we consider the peptide mapping demonstrated. Any mismatches would prompt further investigation.
3. Impurity Profiling by HRMS
3.1 Objective
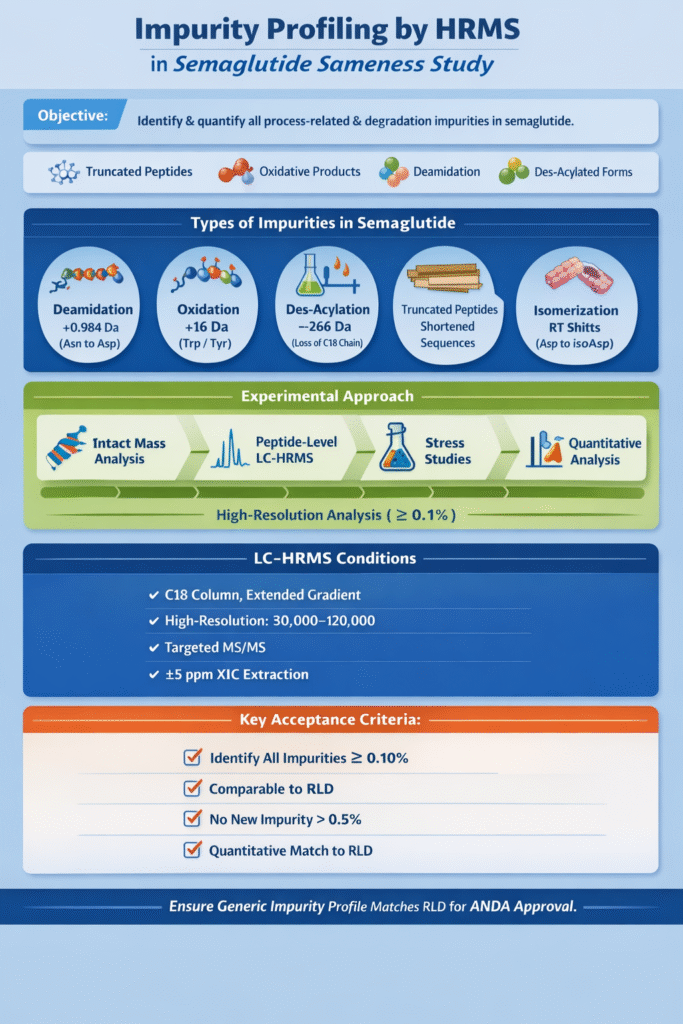
Impurity profiling seeks to identify and quantify all significant process-related or degradation products of semaglutide. Even minor differences in impurity levels can affect safety or efficacy, so regulators require a detailed comparison. We use LC-HRMS to detect variants such as truncated peptides, oxidative or deamidative products, and des-acylated forms.
3.2 Types of Expected Impurities
For a synthetic peptide like semaglutide, common impurities include:
- Deamidation: Asn residues can deamidate to Asp or isoAsp, shifting mass by +0.984 Da per site. (Semaglutide has few Asn, but any present Asn or Gln in the sequence is monitored.)
- Oxidation: Methionine or tryptophan residues (if present) can oxidize (+16 Da). Semaglutide contains a Trp and Tyr, both prone to oxidation.
- Des-acylated semaglutide: Loss of the C18 fatty acid yields a product missing 266 Da. This is a critical impurity to check.
- Truncated peptides: Incomplete synthesis can lead to missing N-terminal or C-terminal residues.
- Isomerization: e.g. Asp formation/isoaspartate can cause retention-time shifts without mass change.
- Aggregation: Although not a small impurity, low-level dimer or aggregate species can form under stress and may be detected by MS or SEC.
Each of these is evaluated. Importantly, peptide-related impurities are excluded from typical ICH Q3A/Q3B rules, so we follow peptide-specific guidance.
3.3 Experimental Approach
Our impurity workflow includes:
- Intact mass scan: First, run the pure peptide (RLD and test) on LC-MS without digestion. The high-resolution full-scan spectrum is checked for additional peaks near the semaglutide mass (e.g. +16, +32, −266 Da, etc). Deconvolution of charge envelopes will reveal any mass variants (oxidized forms, etc).
- Peptide-level analysis: Digest the peptide and run LC-HRMS to pinpoint the location of modifications. For example, an oxidized tryptophan-containing peptide will elute at a shifted time and show +16 Da in its MS.
- Stress studies: We subject semaglutide to forced degradation (oxidative, heat, acid/base) and analyze the degraded samples. This generates likely degradants (e.g. oxidized, deamidated) which helps in identifying them in the stability samples.
- Quantitation: Using extracted ion chromatograms (XICs), we measure peak areas of each impurity relative to the main peak to calculate percentage levels.
All analyses use high-resolution instruments to distinguish isobaric species. For instance, an oxy-semaglutide (+16) and a des-acylated (+x) may have similar m/z in low-res MS, but HRMS resolves them.
3.4 LC-HRMS Conditions
The LC setup for impurity profiling is similar to peptide mapping, with perhaps a longer gradient to separate minor peaks. Key settings include:
- HRMS acquisition: Use full-scan high-resolution MS (e.g. 30,000–120,000 resolving power) over the relevant m/z range.
- Targeted MS/MS: Set up data-dependent or targeted MS/MS for expected impurity m/z (e.g. +16, +266 shifts) to confirm structures.
- XICs: We extract exact-mass chromatograms for suspected impurity formulas (±5 ppm window).
This approach allows detection of low-level impurities, often down to 0.1% of the main peak.
3.5 Data Processing
Data are processed by matching accurate masses and fragmentation patterns to possible structures. In practice, we:
- Use software (e.g. UNIFI, MassHunter) with an accurate-mass screening workflow.
- Search MS/MS spectra against potential impurity masses. For unknowns, we use elemental composition analysis of accurate masses plus observed fragment ions.
- Compare RLD and test impurity maps side-by-side.
Quantification is done by area percentage. We typically report any impurity above the reporting threshold (commonly 0.1% or per ICH Q6B guidelines, which for peptides may be even stricter).
3.6 Key Observations / Acceptance Criteria
For semaglutide, acceptable impurity profiling means:
- Identification of major impurities: All impurities ≥0.10% (w/w) in either RLD or test should be structurally identified.
- Comparability: Any impurity present in the generic should also be present in the RLD at a similar or lower level. In particular, no new peptide-related impurity >0.5% should appear.
- Thresholds: FDA suggests that impurities found in both products at ≥0.1% but ≤0.5% must be justified (e.g. acceptable variation).
- Quantitative match: The generic’s impurity levels should fall within the range observed for the RLD under the same conditions. Significant excess of any impurity may trigger a deficiency.
In summary, the generic’s impurity profile should be comparable to the RLD’s profile. This means that XIC overlays for key masses look nearly identical, and no unexplained peaks appear in the generic. As one guidance slide succinctly states, one must “identify each peptide-related impurity at level ≥0.10%” and ensure no significant new impurity is introduced. Compliant peptide ANDAs present comprehensive HRMS data to meet these criteria.

4. Intact Mass Analysis
4.1 Objective
Intact mass analysis provides an overarching check that the generic semaglutide has the correct molecular weight distribution. It confirms the full peptide mass (with Aib and lipid) and reveals any high-level variants (e.g. mono-oxidized, des-acylated) that may be too low to catch in other assays.
4.2 Method
We inject the unmodified semaglutide (RLD and generic) directly into an LC-MS system under gentle conditions (no digestion). The MS is set for full-scan high-resolution detection. Multiple charge states (typically z = 3–7) are observed in ESI; we deconvolute these to give the neutral masses.
Key notes:
- Deconvolution: Software (e.g. BioPharma finder) collapses the charge envelope to the peptide’s intact mass.
- Resolution: A high resolving power (≥30k) is used so that close variants (e.g. +16, −266) are baseline separated.
4.3 Observations
From intact MS we confirm:
- Correct mass: The expected [M+H]^+ mass of semaglutide (with C18 diacid and Aib) is observed. The deconvoluted mass should match the theoretical mass within <5 ppm.
- Variants: Peaks for minor species appear if present. For example, the monodeamidated (if any), oxidized (+16 Da), or des-acylated (−C18) semaglutide would show up as separate mass peaks. Their identities are confirmed by the exact mass difference.
- Charge-state pattern: The charge distribution of the intact peptide should be identical between samples (any shift could indicate a chemical difference affecting basic sites).
If the intact mass spectra for RLD and generic overlay (no extra peaks) and the main mass matches, this supports sameness. Any unexpected peak here would prompt targeted investigation in peptides or impurities.
5. NMR Characterization (Orthogonal Confirmation)
5.1 Objective
NMR provides an orthogonal structural confirmation that is independent of mass spectrometry. For semaglutide, the key goals of NMR are to verify the peptide backbone and the presence of the lipid moiety. In particular, we use NMR to check that the stearic-acid chain is present and in the correct chemical environment, and that the overall chemical shifts match the reference.
5.2 Techniques
Standard NMR experiments for peptide analysis include:
- 1D ^1H NMR: Gives overall proton fingerprint (amide NH, α-CH, aliphatic CH_2/CH_3 regions).
- 1D ^13C NMR: Identifies carbonyl, aliphatic, and aromatic carbons.
- 2D ^1H–^1H COSY/TOCSY: Reveals proton-proton connectivities within the peptide; useful for assigning spin systems.
- 2D ^1H–^13C HSQC/HMQC: Correlates protons with directly bonded carbons. In peptides, the methyl groups (e.g. of the fatty chain) produce distinct correlations.
- 2D ^1H–^13C HMBC: Long-range C–H correlations (optional).
For semaglutide, high-field NMR (500–700 MHz) may be needed to resolve the densely overlapping peaks. Samples are typically run in DMSO-d6 or a 9:1 H2O/D2O mixture. Acquisition parameters are matched for RLD and generic.
5.3 Key Focus Areas
We pay special attention to:
- Fatty acid chain signals: The stearic diacid has many CH_2 groups and a terminal CH_3. These give characteristic ^1H shifts around 0.8–1.5 ppm and corresponding ^13C shifts. For example, in a 2D ^1H–^13C methyl HSQC, the terminal -CH_3 (at ~0.9 ppm / 14 ppm) should appear. An overlay of the methyl-HMQC spectra between originator and follow-on semaglutide shows matching red/green peaks where identical structure exists.
- Linker (AEEA) region: The 8-amino-3,6-dioxaoctanoic acid spacer has distinct CH_2 and CH signals that can be compared.
- Peptide backbone: The α-protons (4–5 ppm) and amide NH’s (8–9 ppm) of the peptide should match in pattern.
- Chemical shift consistency: Chemical shifts of key signals (e.g. aromatic Tyr, Aib’s α-CH) should be the same in RLD and generic (within ~0.02 ppm), indicating identical local environments.
Crucially, in a 2D overlay of ^1H–^13C spectra, all peaks from the generic should overlay with the reference (no missing or extra peaks).
5.4 Acceptance Criteria
The NMR criteria for semaglutide sameness are:
- Spectral overlap: The NMR spectra of RLD and generic must coincide. Any peak present in one must be present in the other at the same chemical shift.
- Fatty acid confirmation: Signals corresponding to the lipid chain (especially terminal methyl groups) must be present and at the same intensities relative to peptide peaks.
- No extra peaks: Absence of unexpected resonances indicates no chemical impurities.
- Structural fingerprint: Overall, the ensemble of 1D/2D spectra for the generic should serve as a fingerprint indistinguishable from the RLD.
For example, researchers overlaid 2D methyl ^1H–^13C HMQC spectra of originator vs. generic semaglutide and found that all methyl resonances (green circles in the figure) matched. A comparable result here would strongly confirm structural identity.
6. Comparative Sameness Assessment (RLD vs Test)
After gathering all data, we compile a comparative assessment. This integrates the results from sequence confirmation, peptide mapping, impurity profiling, intact mass, and NMR. The generic is considered the “same” as the RLD if:
- Sequence and structure: The amino acid sequence (including Aib) and post-synthetic modifications match exactly (MS/MS results).
- Peptide map: Chromatograms of digested peptides from RLD and generic overlay perfectly (no unmatched peaks).
- Impurity profile: All significant impurities in the generic were present in the RLD at similar levels (and have been identified).
- Intact mass: The intact molecular weight and charge-state pattern are identical, with any minor variants (e.g. oxidized, des-acylated species) being comparable in both samples.
- NMR fingerprint: (If performed) The NMR spectra of RLD and generic coincide completely, confirming the same chemical structure.
If all these orthogonal data align, one can statistically or qualitatively assert molecular identity. As ResolveMass emphasizes, combining these techniques into “a coherent comparability narrative” is essential. Any minor discrepancies (e.g. low-level impurity) must be scientifically justified with reference to the RLD data. When done thoroughly, the multi-technique approach leaves little doubt about sameness.
7. Regulatory Considerations for ANDA
From a regulatory perspective, demonstrating semaglutide sameness involves both formulation sameness and structural equivalence. For products like semaglutide injection, FDA guidance stipulates Q1/Q2 sameness of excipients for a bioequivalence waiver. This means the generic’s formulation must use the same ingredients at the same concentrations (±5%) as the RLD.
At the API level, however, the standards are even stricter. The generic semaglutide must be shown to be molecularly identical to the RLD, per FDA’s ANDA peptide guidance. The FDA explicitly expects: “Exact amino acid sequence, same peptide length, no substitutions, deletions, insertions or unintended modifications”. As noted in FDA training slides, peptide API sameness is established by matching the primary sequence, physicochemical properties, aggregation behavior, and biological activity. The comprehensive analytical package we’ve outlined meets these requirements by using multiple orthogonal methods for each attribute.
Peptide APIs are evaluated under the drug (505(j)) framework, not biosimilar rules, meaning identity, not just similarity, is required. Thus, data from LC-HRMS, LC-MS/MS, and NMR must definitively show no meaningful differences. FDA and ICH guidance underscore the need for “state-of-the-art” analytical tools and 100% sequence coverage. Indeed, Waters notes that an LC-HRMS workflow can provide “qualitative and quantitative analysis of synthetic peptide impurities…in accordance with ICH and FDA guidelines”.
In sum, the regulatory strategy for a semaglutide ANDA is to present a robust comparability dossier: confirming Q1/Q2 for the formulation and providing peer-reviewed-quality structural data for the API. This reduces regulatory risk, as deficiencies in peptide characterization are a common cause of review delays.
Conclusion:
A Semaglutide Sameness Study is a multi-faceted analytical endeavor that underpins generic approval. It requires precise amino acid sequencing, exhaustive impurity comparability, and confirmation of structural fidelity through orthogonal methods. When done correctly, it satisfies FDA requirements and enables waivers of unnecessary studies (like in vivo trials) for semaglutide generics. Importantly, early investment in a comprehensive peptide sameness package pays off by minimizing review issues and expediting approval.
ResolveMass Laboratories is ready to guide you through this process. Our peptide analytical experts will design a tailored sameness study, execute it with high precision, and interpret the results in the context of FDA ANDA guidance. We provide clear, defensible reports so that regulators have full confidence in your ANDA submission.
Frequently Asked Questions:
A Semaglutide Sameness Study is a comprehensive analytical evaluation performed to demonstrate that a generic version of semaglutide is structurally and functionally equivalent to the reference listed drug (RLD), using techniques like LC-MS, HRMS, peptide mapping, and NMR.
Peptide mapping provides a comparative fingerprint of the molecule, confirming that the generic and RLD produce identical peptide profiles with no missing or additional peaks.
Key techniques include:
-HRMS (High-Resolution Mass Spectrometry)
-LC-MS/MS (Liquid Chromatography–Mass Spectrometry)
-Peptide Mapping
-Enzymatic Digestion (Trypsin, Glu-C, etc.)
-1D & 2D NMR (COSY, HSQC)
These orthogonal methods ensure complete structural confirmation.
Common impurities include:
-Deamidation (+0.984 Da)
-Oxidation (+16 Da)
-Des-acylated forms (−266 Da)
-Truncated peptides
-Isomerization (Asp/isoAsp)
These must be identified and quantified during impurity profiling.
HRMS enables accurate mass measurement and high-resolution detection of impurities, allowing identification of low-level variants and differentiation between closely related molecular species.
Reference
- Kim SH, Kim SS, Kim HJ, Park EJ, Na DH. Peptide mapping analysis of synthetic semaglutide and liraglutide for generic development of drugs originating from recombinant DNA technology. Journal of Pharmaceutical and Biomedical Analysis. 2025 Apr 15;256:116682.https://www.sciencedirect.com/science/article/pii/S0731708525000238
- De Groot AS, Mattei A, Gabriel B, Calderini J, Roberts BJ, Lelias S, McAllister M, Boyle C, Martin W, Richard G. Immunogenicity of generic peptide impurities: current orthogonal approaches. Pharmaceutical research. 2025 May;42(5):805-18.https://link.springer.com/article/10.1007/s11095-025-03843-1
- Rogers-Crovak JA, Delaney EJ, Detlefsen DJ. Recommendation for Clarifying FDA Policy in Evaluating “Sameness” of Higher Order Structure for Generic Peptide Therapeutics. The AAPS Journal. 2024 Nov 26;27(1):8.https://link.springer.com/article/10.1208/s12248-024-00994-8
- Giri T, Sakharwade S, Subbappa P, Chinnakadoori SR, Sharma N. Regulatory Considerations in Synthetic Peptide Characterization: Techniques and Compliance. Separation Science Plus. 2025 Jun;8(6):e70057.https://analyticalsciencejournals.onlinelibrary.wiley.com/doi/abs/10.1002/sscp.70057


