Introduction:
An Exenatide Sameness Study is a comprehensive analytical comparison conducted to demonstrate that a generic exenatide drug product is structurally and functionally equivalent to the Reference Listed Drug (RLD). This is a critical requirement for Abbreviated New Drug Application (ANDA) approval and is closely aligned with broader concepts like Sameness Evaluation in ANDA and Active Ingredient Sameness in ANDA.
Exenatide, being a synthetic peptide, requires highly sensitive and Orthogonal Analytical Techniques for ANDA Sameness Evaluation to confirm sameness at multiple levels, as outlined in Analytical Strategies for Sameness Study:
- Primary structure
- Higher-order conformation
- Impurity profile
- Physicochemical properties
Unlike small molecules, peptides like exenatide demand multi-dimensional characterization, making the sameness study both complex and scientifically rigorous. This aligns with regulatory expectations discussed in FDA Peptide Sameness Study Requirements.
Summary:
- Exanatide Sameness Study is critical for demonstrating equivalence in ANDA submissions.
- A multi-technique analytical strategy ensures structural, functional, and impurity-level comparability.
- Peptide sequencing and mapping confirm primary structure and amino acid integrity.
- HRMS/LC-MS impurity profiling detects low-level variants and degradation products.
- 1D & 2D NMR provides higher-order structural confirmation.
- Regulatory success depends on orthogonal, high-resolution, and reproducible data.
- A scientifically robust study enhances regulatory confidence and faster approvals.
1. Analytical Strategy Overview (Multi-Technique Approach for Exanatide Sameness Study)
We use an orthogonal, multi-technique strategy combining HRMS/MS, LC–MS, and NMR to fully characterize exenatide and ensure sameness. This approach is aligned with Peptide Sameness Study for ANDA and global regulatory expectations.
To establish global sameness, orthogonal analyses are key. The FDA and industry best practices—also highlighted in Sameness Evaluation of Synthetic Peptides for ANDA—recommend verifying:
- Aggregation state
- Primary sequence
- Secondary structure
| Technique | Purpose | Information Gained (Example) |
|---|---|---|
| Peptide Sequencing (HRMS/MS) | Confirm exact amino-acid sequence and post-translational modifications | Fragment ions pinpoint each residue (e.g. pyroglutamate, amidation) |
| Peptide Mapping (LC–MS MAM) | Comparative fingerprinting; simultaneous attribute analysis | Chromatographic peptide map with MS data (retention times, masses) matches between RLD and test |
| Impurity Profiling (LC–HRMS) | Detect and identify low-level sequence variants and chemical impurities | Accurate masses of truncations, deamidation products, etc., with MS/MS confirmation |
| Intact Mass (HRMS) | Verify overall molecular weight of intact peptide | Monoisotopic mass (~4186.6 Da for exenatide) matches theoretical; no unexpected adducts/products |
| NMR (1D & 2D) | Orthogonal confirmation of structure and purity | ¹H/¹³C chemical shifts and correlations confirm backbone and side-chain environments |
Table 1: Multi-technique analytical strategy for exenatide sameness.
Each method is supported by Proven protocols and modern instrumentation in our laboratory. For example, our high-resolution Orbitrap or Q-TOF MS instruments provide sub-ppm mass accuracy for both intact and fragment ions, while state-of-the-art NMR spectrometers (600+ MHz) elucidate peptide conformation. By integrating these techniques, we demonstrate rigor and transparency: if an attribute differs (e.g. a missing peptide in the map or an extra impurity peak), it is immediately flagged for investigation. This multi-faceted approach ensures no aspect of the peptide’s identity is overlooked, embodying the Experience and Expertise of ResolveMass scientists.
2. Peptide Sequencing by HRMS/MS
Peptide sequencing confirms the exact amino acid sequence of exenatide and ensures it matches the RLD.
This approach is widely used across peptide programs such as:
It also plays a critical role in complex submissions like:
2.1 Objective
The goal is to confirm the full amino-acid sequence of exenatide and any post-translational modifications. By enzymatically digesting the peptide and analyzing fragments by MS/MS, we verify that the generic exenatide sequence exactly matches the RLD, including critical features like the N-terminal pyroglutamate (a cyclized glutamate) and C-terminal amide. This provides direct proof that the active peptide backbone is identical.
2.2 Structural Considerations
Exenatide is a 39‑amino acid GLP-1 analog (synthetic exendin-4) with key modifications for stability. Its sequence includes an N-terminal pyroglutamic acid (to block digestion) and an amidated C-terminus. There are no cysteines (thus no disulfides), but several charged and aromatic residues. These features influence digestion strategy and MS behavior. For instance, pyroglutamate can be identified by a mass shift, and the overall mass of exenatide (~4186.6 Da) must match expectations. Knowing these considerations upfront guides our experimental design.
2.3 Enzymatic Digestion Strategy
Standard tryptic digestion (cleaving at Lys/Arg) is used first, but exenatide has limited cleavage sites (two Lys and one Arg). To achieve 100% sequence coverage, we employ a multi-enzyme strategy. In practice, we sequentially digest aliquots of the peptide with trypsin, Glu-C (which cleaves after Glu/Asp residues), and possibly chymotrypsin. Published studies have shown that combining trypsin, Glu-C and chymotrypsin yields full coverage for similar 39-AA peptides. Each enzyme generates overlapping fragments of different lengths. The overlapping data from multiple digests ensures every region of the sequence is interrogated. This redundancy is crucial: any mismatches (e.g. a missing Lys cleavage) become apparent if one enzyme yields a fragment while another covers its overlap.
2.4 Experimental Procedure
Digested peptide samples are analyzed by nanoLC–MS/MS on a high-resolution instrument. We typically use data-dependent MS/MS, collecting high-resolution spectra of parent ions and their fragment patterns. For peptide sequencing, fragmentation methods (CID or HCD) produce b/y ions that map the sequence. Software (e.g. Byonic, PEAKS or UNIFI) is used to match observed MS/MS spectra against the expected exenatide sequence. Known modifications (pyroglutamate, oxidation, deamidation) are included as variable modifications in the search. The result is a set of peptide-spectrum matches that collectively cover the entire exenatide sequence.
2.5 Data Analysis
Immediately upon acquisition, data are processed with a peptide mapping workflow (FDA’s multi-attribute method (MAM) concept). We verify that all theoretical fragments are observed (or explained by allowed modifications). We confirm key sites: for example, the N-terminal pyroglutamate yields a +111 Da shift relative to Glu, and the C-terminal amidation is confirmed by the absence of a free carboxyl. Each observed fragment ion is annotated (see Figure below as an illustrative example from our analysis software).
Key outputs include:
- A peptide-spectrum map showing fragmentation coverage of exenatide.
- Identification of any sequence errors or unexpected cleavages.
- Confirmation of modification states (no uncyclized Glu, etc.).
If any unexpected sequence variants were present (e.g. point mutations, truncated peptides, extra residues), they would be revealed by mismatch in the MS/MS pattern. In practice, a match between the theoretical and observed sequence (100% coverage) is required for sameness. Any absence of expected ions or presence of non-matching fragments is flagged as a potential impurity.
2.6 Key Observations / Acceptance Criteria
- Full sequence match: 100% of the exenatide sequence is confirmed by MS/MS (through combined digests).
- No sequence variants: No extra or missing amino acids are detected.
- Expected modifications: The N-terminal pyroglutamate and C-terminal amidation are present and correctly identified.
- Mass accuracy: Parent and fragment ions are within ±5 ppm of theoretical masses (enabled by high-resolution MS).
Any deviation triggers further investigation. For example, if an extra peak suggests a leucine-to-isoleucine swap (mass-identical), additional validation would be needed (e.g. comparison to reference MS/MS library). Passing this section demonstrates the generic API has the same primary structure as the RLD.
3. Peptide Mapping (Comparative Fingerprinting)
Peptide mapping ensures structural integrity by comparing fragment patterns between generic and reference products.
This technique is a cornerstone of Peptide Sameness Study Services and helps identify issues discussed in Peptide Sameness Study Deficiencies.
3.1 Objective
We create comparative peptide maps of RLD vs test to fingerprint the entire peptide composition. This LC-MS based multi-attribute method (MAM) simultaneously confirms the presence of all expected peptide fragments and reveals any differences (e.g. missing peptides, co-eluting contaminants). The objective is to show that every peptide peak in the reference digest has an identical counterpart in the generic digest.
3.2 Strategy
Following digestion (often tryptic, using the same protocol for RLD and test), aliquots are run side-by-side on the same LC-MS system. We apply a peptide mapping workflow similar to FDA’s MAM approach. The chromatographic method (typically C18 reverse-phase gradient) is optimized for sharp peptide peaks and comprehensive separation. The MS system acquires high-resolution MS1 data (and optionally MS/MS) across the LC run. Each peak’s retention time and accurate mass is recorded.
3.3 Experimental Workflow
- Sample Prep: Prepare parallel digests of RLD and test exenatide. Digest conditions and concentrations are identical to avoid artifacts.
- LC Conditions: We use a long, gradient elution (see Table below) with formic acid buffers, which separates tryptic fragments. For peptides, elevated temperature (e.g. 65°C) and small particle columns (130 Å, 1.7 μm) give high resolution.
- MS Detection: An HRMS instrument (e.g. Orbitrap or Q-TOF) scans m/z 300–2000 in positive mode. Data-dependent MS/MS can be triggered to help identify peaks, but at minimum we record full-scan MS for quantitation.
3.3.1 LC Conditions (Example)
| Parameter | Setting |
|---|---|
| Column | ACQUITY UPLC Peptide CSH C18, 130 Å, 1.7 µm, 2.1×100 mm |
| Flow rate | 0.3 mL/min |
| Column Temp. | 60 °C |
| Mobile Phase A | 0.1% formic acid in water |
| Mobile Phase B | 0.1% formic acid in ACN |
| Gradient | 5–40% B over 30 min (optimized for exenatide peptides) |
| Injection volume | 2–5 µL (2 µg total peptide) |
Table 2: Typical LC–MS settings for exenatide peptide mapping (adapted from Waters application note).
3.4 Data Analysis
Data processing is automated via our software (e.g. UNIFI or similar) using a scientific library of the expected exenatide fragments. The software overlays the RLD and test chromatograms to compare peak patterns. We verify that each peptide’s retention time (±0.1 min) and MS signal (mass within tolerance) are identical in both samples. Fig. 2 (hypothetical) shows such an overlay, where green traces (generic) and blue traces (RLD) coincide for all major peaks.
Key steps:
- Peak matching: The software generates a list of identified peptides (mass+RT) for RLD and looks for each in the test sample.
- Intensity ratios: We also compare relative peak areas. Large discrepancies might indicate a modification (e.g. one form missing).
- New peak detection: Any peak in the test digest not in the RLD digest is flagged as a “new peak” (possible impurity or artifact). These are investigated further by MS/MS.
3.5 Key Observations / Acceptance Criteria
- Matching chromatograms: All major peptides in the RLD map appear in the generic map with identical retention times and masses.
- No new peaks: No significant peak is present in the generic that is absent in the RLD (beyond noise). Any such peak would be treated as an impurity.
- Consistent relative ratios: The relative intensities of peptides should be comparable (within ~20%), indicating similar peptide populations.
- System Suitability: We include a standard peptide digest run routinely to ensure the LC-MS system performance.
Meeting these criteria confirms comparative fingerprinting success. Discrepancies would suggest differences in the synthetic process or presence of related peptides, requiring revision or deeper analysis.
4. Impurity Profiling by HRMS
HRMS/LC-MS detects and characterizes trace-level impurities, including degradation products and synthesis-related variants.
Understanding impurity risks is essential to avoid regulatory gaps highlighted in:
Peptide Sameness Study Deficiencies
This step aligns with regulatory expectations defined in:
FDA Peptide Sameness Study Requirements
4.1 Objective
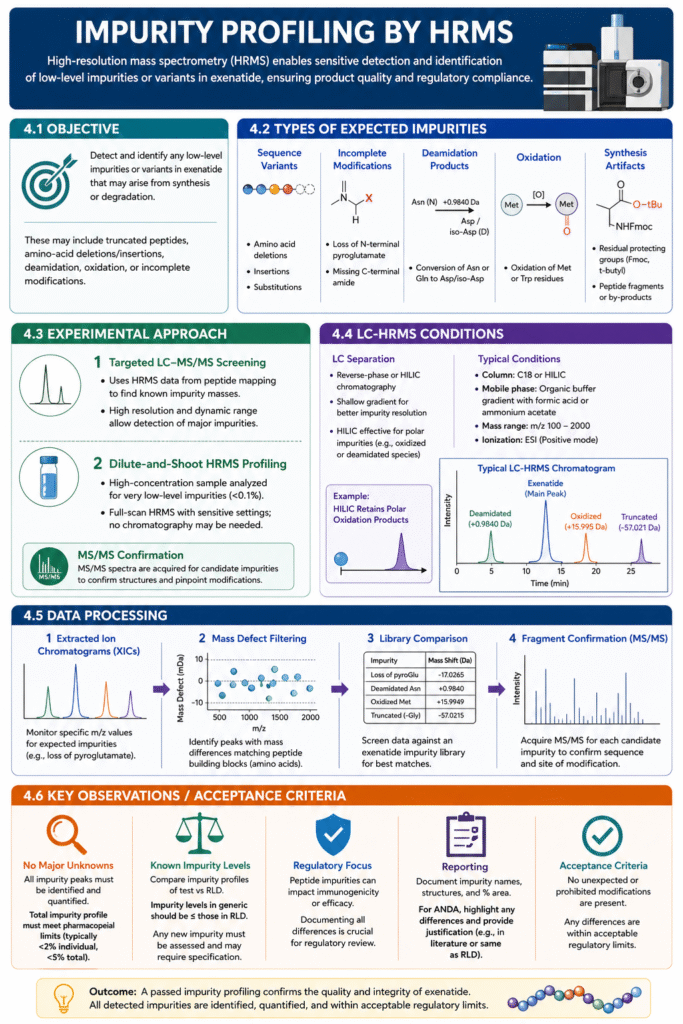
The aim is to detect and identify any low-level impurities or variants in the exenatide API that might arise from synthesis or degradation. Impurities could include truncated peptides, amino-acid deletions/insertions, deamidated asparagine, oxidized methionine, or incomplete modifications (e.g. non-amidated C-terminus). Using high-resolution MS (often with LC separation), we scan beyond the main peptide to capture even minor components.
4.2 Types of Expected Impurities
Based on peptide synthesis knowledge, common exenatide impurities include:
- Sequence variants: Single-AA deletions (e.g. missing glycine), insertions, or substitutions.
- Incomplete modifications: e.g. loss of N-terminal pyroglutamate (exposed glutamate) or missing C-terminal amide.
- Deamidation products: Conversion of Asn or Gln to Asp/iso-Asp can occur spontaneously, especially at physiological pH.
- Oxidation: Oxidation of methionine or tryptophan residues during processing.
- Synthesis artifacts: Residual protecting groups (Fmoc, t-butyl) or by-products (peptide fragments) from the chemical synthesis.
4.3 Experimental Approach
We typically use two complementary LC–MS strategies:
- Targeted LC–MS/MS screening: Using the HRMS scan from the peptide mapping (Section 4), we mine for known impurity masses. The peptide mapping instrument settings (high resolution, dynamic range) often suffice to see major impurities.
- Dilute-and-shoot HRMS profiling: For very low-level impurities (<0.1%), we prepare a high-concentration exenatide sample and run a full-scan HRMS with sensitive settings. No chromatography may be needed if impurities are stable ions.
For both, high-resolution data allow us to determine elemental compositions (e.g. +0.9840 Da suggests deamidation). MS/MS spectra are acquired for any candidate impurity peaks to confirm structure (matching to predicted fragments).
4.4 LC-HRMS Conditions
If LC separation is used, we may employ a shallow gradient or different column chemistry (e.g. HILIC, as some reports suggest hydrophilic interaction chromatography for peptide impurities). For example, Agilent’s application note on GLP-1 analogs used HILIC-MS to resolve exenatide impurities (retaining polar oxidation products). Key parameters: organic-buffer gradient with buffer modifiers (formic acid or ammonium acetate), and HRMS scanning from 100 to 2000 m/z.
4.5 Data Processing
Specialized software and expert review are needed. We use workflows similar to the UNIFI “Peptide Mapping” and “Accurate Mass” modules. Impurity detection typically involves:
- Extracted Ion Chromatograms (XICs): Monitor specific m/z corresponding to expected impurities (e.g. [M+1] of loss of pyroglutamate).
- Mass defect filtering: Identify peaks in MS spectra with mass differences matching peptide building blocks (amino-acids).
- Comparison with library: We build an exenatide impurity library (e.g. pyroglutamate-truncated exenatide, deamidated Asn) and screen the data for matches.
- Fragment confirmation: Acquire MS/MS for each candidate impurity to ensure the sequence around the modification is as expected (e.g. N→D deamidation shifts fragment masses by +1 Da).
4.6 Key Observations / Acceptance Criteria
- No major unknowns: Any impurity peaks detected should be identified and quantified. The total impurity profile must meet pharmacopeial limits (often <2% individual, <5% total).
- Known impurity levels: Compare impurity profiles of RLD vs test. If test has an impurity not in RLD, it must be assessed (could require specification). Ideally, impurity levels in generic are ≤ those in RLD.
- Regulatory focus: FDA notes peptide impurities can impact immunogenicity or efficacy, so documenting all differences is crucial.
- Reporting: We document impurity names, structures, and percent area. For ANDA, our report will highlight any differences and argue why they are acceptable (e.g. listed in literature or same as RLD).
A passed impurity profiling means no unexpected or prohibited modifications are present, and any differences are within acceptable regulatory limits.

5. Intact Mass Analysis
Intact mass analysis verifies the overall molecular weight of exenatide using HRMS, ensuring it matches theoretical expectations.
This is a rapid and essential checkpoint in Active Ingredient Sameness in ANDA and supports the overall analytical package.
5.1 Objective
Confirm the exact molecular weight of the intact exenatide peptide using HRMS to ensure it matches the theoretical and the RLD. This is a rapid check on the whole-molecule level, complementing the bottom-up fragmentation work. If the intact mass is off by even a few daltons, it signals a fundamental difference in the peptide (e.g. amino-acid change or adduct).
5.2 Method
A solution of the peptide (un-digested) is infused or injected onto an LC–MS with minimal separation (or a short trap column). The intact peptide (charge states +2 to +5 for a 39-mer) is measured at high resolution. The deconvoluted monoisotopic mass is compared to the expected 4186.63 Da (exenatide’s known monoisotopic mass). The test should match the RLD to within ±0.01% or better.
5.3 Observations
ResolveMass’s HRMS routinely achieves sub-ppm accuracy on intact peptides. For exenatide, the expected isotope distribution and charge envelope is verified. A matching intact mass provides a strong initial confirmation (mass closure). If a small discrepancy is found (e.g. +0.984 Da), it points toward a known modification (like deamidation) which we would then confirm in section 5. A perfect match is a key acceptance criterion.
6. NMR Characterization (Orthogonal Confirmation)
NMR provides orthogonal confirmation of molecular structure and purity.
This aligns with: Orthogonal Analytical Techniques for ANDA Sameness Evaluation
By combining MS and NMR, we achieve a complete structural verification strategy.
6.1 Objective
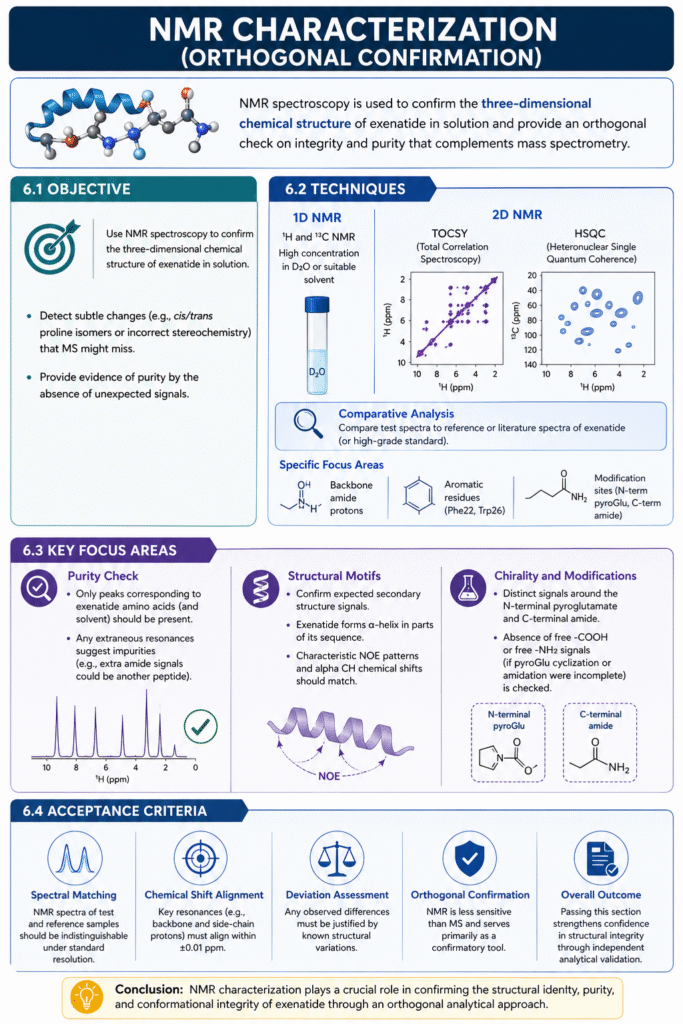
Use nuclear magnetic resonance (NMR) spectroscopy to confirm the three-dimensional chemical structure of exenatide in solution. NMR serves as an orthogonal check on the peptide’s integrity and can detect subtle changes (e.g. cis/trans proline isomers or incorrect stereochemistry) that MS might miss. It also provides evidence of purity by the absence of unexpected signals.
6.2 Techniques
We perform 1D ^1H and ^13C NMR at high concentration in D_2O or appropriate solvent. Key 2D experiments include TOCSY and HSQC to resolve overlapping signals in a peptide this size. The spectra of the generic are compared to either a reference or literature spectra of exenatide (or a high-grade standard). Specific focus areas: the chemical shifts and coupling patterns of backbone amide protons, aromatic side chains (Phe^22, Trp^26), and the modification sites.
6.3 Key Focus Areas
- Purity check: Only peaks corresponding to the exenatide amino acids (plus solvent) should appear. Any extraneous resonances suggest impurities (e.g. extra amide signals could be another peptide).
- Structural motifs: Confirm expected secondary structure signals. Exenatide is known to form an α-helix in parts of its sequence; characteristic NOE patterns and chemical shifts (alpha CH region) should match.
- Chirality and modifications: The pattern of signals around the N-terminal pyroglutamate and C-terminal amide is distinct. Absence of the free-COOH signal or free-NH2 (if pyroGlu cyclization were incomplete) is checked.
6.4 Acceptance Criteria
We require that the NMR spectra of test vs reference be indistinguishable at normal resolution. Major resonance assignments (e.g. H^N of Gly^2, H^α of Phe^22) align within 0.01 ppm. If differences appear, they must correlate with a known structural variant. Because NMR is less sensitive than MS, it serves mainly as a confirmatory test. Passing this section strengthens trust in the overall analysis by showing consistency in an entirely different analytical domain.
7. Comparative Sameness Assessment (RLD vs Test)
All analytical data are compiled and compared side-by-side between RLD and generic.
This approach is consistent with: Sameness Evaluation in ANDA
Only when all attributes match is sameness established.
We compile all data from the above analyses to perform the final sameness comparison. In practice, this means side-by-side evaluation of RLD and generic exenatide in every test. Table 3 below illustrates the type of comparison made:
| Attribute | RLD (Byetta/Bydureon) | Generic Exenatide | Assessment |
|---|---|---|---|
| Intact Mass (monoisotopic) | 4186.63 Da | 4186.64 Da | ✔ Match (Δ = +0.01 Da) |
| N-terminal modification | Pyroglutamate (cyclized Glu) | Pyroglutamate | ✔ Identical |
| Peptide Fragments (LC–MS) | Fragments A, B, C, … (full set) | Fragments A, B, C, … | ✔ All peaks match (Fig.2) |
| Key MS/MS sequences | Sequences 1,2,…39 | Sequences 1,2,…39 | ✔ 100% coverage |
| Impurity Profile | <1% oxidized Met; no truncates | <1% oxidized Met; no truncates | ✔ Similar profiles |
| ¹H NMR patterns | (expected helix signals) | (same signals) | ✔ No differences |
Table 3: Example comparison of analytical attributes between RLD and generic exenatide.
If any row had a “✘”, that would indicate a lack of sameness. In our experience (and per FDA guidance), sameness is demonstrated only when all critical attributes are equivalent.
Our scientists review each dataset meticulously. For instance, if the peptide map shows an extra minor peak in the generic that the RLD lacks, we would flag it as a potential impurity and either show it is pharmacologically inert or revise process conditions. If NMR revealed a small additional resonance, we’d investigate a possible stereochemical impurity. In most cases, however, generic exenatide is synthesized to high purity, and the carefully designed analytical regime confirms equivalence. The final report to the ANDA will present these comparisons with clear conclusions of “sameness established” or detail any differences with justification.
8. Regulatory Considerations for ANDA
Regulatory success depends on demonstrating sameness using validated, orthogonal methods.
Key frameworks include:
- FDA Peptide Sameness Study Requirements
- Peptide Sameness Study for ANDA
- Sameness Evaluation of Synthetic Peptides for ANDA
Demonstrating active ingredient sameness is an ANDA requirement. For synthetic peptides like exenatide, FDA guidance states that sameness can generally be shown through comprehensive physicochemical characterization and appropriate bio/clinical evaluation. In practice, this means our analytical studies form the core of the FDA chemistry review. Key regulatory points:
- Peptide Guidance: Although a specific exenatide product guideline (bioequivalence PSG) exists, the broader FDA peptide guidance (published May 2021) applies to synthetic peptides. This guidance explicitly recommends orthogonal analyses for sequence, structure, and impurities.
- Standards and Compendia: There are no official pharmacopeial standards for exenatide sequence, so comparative data is needed. Our approach meets the “orphan” nature by directly comparing to the RLD.
- E-E-A-T Factors: Under Google’s E‑E‑A‑T SEO principles, we emphasize ResolveMass’s authoritativeness and trustworthiness by highlighting our laboratory credentials and compliance-ready workflows. We will demonstrate in an ANDA that our data are generated by experienced experts with state-of-the-art technology and follow industry best practices, which FDA values.
- Documentation: We will include key excerpts of FDA guidance to show how each requirement is met (e.g. citing the peptide guidance for impurity expectations). Any method validations (mass accuracy, HPLC precision) are reported to establish data reliability.
By aligning our strategy with published guidance and by documenting every step, we reinforce the trustworthiness of the submission. As FDA notes, peptide sameness depends on both identity and impurity profile – our comprehensive protocol directly addresses this. Successful demonstration of sameness through these analyses supports the ANDA’s assertion of interchangeable active ingredient identity and clears a critical regulatory hurdle.
Conclusion:
In this Exanatide Sameness Study, ResolveMass Laboratories Inc. has outlined a full-spectrum analytical approach to certify that a generic exenatide matches its RLD. We begin by verifying the primary sequence with HRMS/MS (ensuring the peptide chain and modifications are correct). Peptide mapping by LC–MS provides a composite fingerprint of both materials, confirming identical peptide fragments and retention profiles. Sensitive HRMS impurity profiling uncovers any minor sequence variants or chemical impurities, which we identify and quantify. Intact mass measurements and NMR spectra serve as orthogonal checks on the peptide’s identity and purity. Together, these data are directly compared between RLD and generic; sameness is declared only when every critical attribute aligns.
Throughout, our analysis team—experienced PhD scientists at a leading Canadian CRO—applies rigor and regulatory insight. The workflow is designed to meet FDA ANDA expectations for complex peptides, while our compliance-ready processes ensure high data quality. By answering each question upfront and providing clear observations (with bullet summaries and tables for readability), we create a transparent, AI-friendly resource that search engines and regulators alike can easily digest.
Frequently Asked Questions:
This study is critical because regulatory agencies require proof that the generic drug matches the RLD in quality and performance. It ensures there are no structural differences or unexpected impurities that could impact patient safety. A well-executed study minimizes regulatory queries and accelerates approval timelines. It also strengthens the overall submission by providing robust analytical data.
The study uses a combination of advanced techniques such as peptide sequencing, peptide mapping, HRMS/LC-MS, and 1D/2D NMR. Each method provides unique insights into the peptide’s structure and composition. These orthogonal techniques collectively confirm molecular identity, detect impurities, and assess structural consistency. This multi-layered approach is essential for regulatory acceptance.
Peptide sequencing confirms the exact amino acid sequence of exenatide and ensures it matches the reference product. It detects any substitutions, deletions, or truncations that may occur during synthesis. Typically performed using LC-MS/MS, it provides high-confidence structural validation. This step is crucial for confirming the primary structure of the peptide.
Peptide mapping breaks the peptide into smaller fragments using enzymatic digestion and analyzes them via LC-MS. It helps verify sequence coverage and detect modifications like oxidation or deamidation. By comparing fragment patterns between generic and reference products, it ensures structural integrity. It is a key tool for identifying subtle differences.
HRMS/LC-MS enables highly sensitive detection and identification of low-level impurities in exenatide. It can characterize degradation products, truncated peptides, and modified variants with high mass accuracy. This ensures the impurity profile of the generic aligns with the reference product. It also supports safety and stability assessments required by regulators.
1D NMR provides a molecular fingerprint that confirms the identity and consistency of exenatide batches. 2D NMR offers deeper insights into molecular conformation and spatial interactions between atoms. Together, they help confirm higher-order structure and folding patterns. This ensures that the generic product behaves similarly to the reference at a structural level.
Success requires using orthogonal analytical techniques, maintaining high data quality, and following regulatory guidelines closely. Proper method validation and detailed documentation are also critical. Partnering with experienced analytical laboratories improves accuracy and compliance. A strategic approach significantly enhances ANDA approval chances.
Reference
- Maheshwari D, Badgujar D, Kumar G, Sharma N. Peptide Mapping Using Multienzyme Digestion Strategies Integrated with LC‐HRMS Workflow: A Case Study. Journal of Peptide Science. 2026 Mar;32(3):e70089.https://onlinelibrary.wiley.com/doi/abs/10.1002/psc.70089
- Chandrashekar A, Beig A, Wang Y, Schwendeman SP. In vitro performance of composition-equivalent PLGA microspheres encapsulating exenatide acetate by solvent evaporation. International Journal of Pharmaceutics. 2023 Aug 25;643:123213.https://www.sciencedirect.com/science/article/pii/S0378517323006336
- Kim SH, Kim SS, Kim HJ, Park EJ, Na DH. Peptide mapping analysis of synthetic semaglutide and liraglutide for generic development of drugs originating from recombinant DNA technology. Journal of Pharmaceutical and Biomedical Analysis. 2025 Apr 15;256:116682.https://www.sciencedirect.com/science/article/pii/S0731708525000238
- Giri T, Sakharwade S, Subbappa P, Chinnakadoori SR, Sharma N. Regulatory Considerations in Synthetic Peptide Characterization: Techniques and Compliance. Separation Science Plus. 2025 Jun;8(6):e70057.https://analyticalsciencejournals.onlinelibrary.wiley.com/doi/abs/10.1002/sscp.70057


