Summary:
Dulaglutide Sameness Study is a comprehensive analytical comparison demonstrating that a proposed generic dulaglutide product is structurally and chemically identical to the reference listed drug (RLD).
An orthogonal, multi-technique approach is used: peptide sequencing and mapping (via LC-MS/MS) verify the exact amino acid sequence and coverage, HRMS/LC-MS profiling detects trace impurities and variants, and 1D/2D NMR spectroscopy confirms higher-order structure.
- Similar approaches are used in Liraglutide Sameness Study and Semaglutide Sameness Study
- Regulatory gaps can be avoided by understanding Peptide Sameness Study Deficiencies
- Compliance is guided by FDA Peptide Sameness Study Requirements
For example, tryptic peptide mapping of dulaglutide achieved ~93% sequence coverage, ensuring nearly complete verification of its primary structure.
High-resolution LC-MS (HRMS) enables detection and identification of low-level process- or degradation-related impurities.
1D/2D NMR provides orthogonal confirmation of dulaglutide’s chemical structure.
- Services are available globally via Peptide Sameness Study Services in United States and Peptide Sameness Study Services in Canada
1. Introduction:
Proving structural sameness is critical for ANDA approval. The Dulaglutide Sameness Study ensures identity, purity, and higher-order structure match the RLD.
- This aligns with broader concepts like Peptide Sameness Study for ANDA
- Similar case studies include Octreotide Sameness Study for ANDA Submission and Liraglutide Sameness Study for ANDA Submission
Regulators emphasize orthogonal analysis, ensuring complete comparability and reducing ANDA deficiencies.
2. Analytical Strategy Overview (Multi-Technique Approach)
A Dulaglutide Sameness Study relies on orthogonal analytical techniques to confirm equivalence.
- This aligns with Analytical Strategies for Sameness Study
- And Orthogonal Analytical Techniques for ANDA Sameness Evaluation
| Technique | Purpose | Information Gained |
|---|---|---|
| Peptide Sequencing (HRMS/MS) | Confirm exact amino-acid sequence and modifications | Fragmentation (b/y ions) identifies each residue and PTMs |
| Peptide Mapping (LC–MS/MAM) | Comparative fingerprint of digested peptides | LC retention time + mass of each peptide matches RLD map |
| Impurity Profiling (LC–HRMS) | Detect and identify low-level impurities/variants | Accurate mass of truncations, deamidations, oxidations; MS/MS confirm structure |
| Intact Mass (HRMS) | Verify overall molecular weight | Monoisotopic mass (~59,669 Da for dulaglutide) matches theory |
| NMR (1D & 2D) | Orthogonal structure confirmation | ¹H/¹³C chemical shifts and 2D correlations confirm folding and purity |
- Similar peptide comparisons are discussed in Sameness Evaluation of Synthetic Peptides for ANDA
This multi-faceted plan ensures that any discrepancy – be it a missed amino acid, an extra modification, or an unexpected impurity – will be caught. By covering sequence, composition, mass and structure in parallel, we build a complete, defensible sameness package.
3. Peptide Sequencing by HRMS/MS
3.1 Objective
The goal of peptide sequencing is to confirm the entire primary structure of dulaglutide at the amino-acid level. We must verify 100% sequence identity with the reference (no substitutions, deletions or insertions). Any difference would signal a fundamentally different molecule. Thus, we fragment the peptide in MS/MS and match the observed fragments to the expected dulaglutide sequence, ensuring perfect concordance.
3.2 Structural Considerations
Dulaglutide is a 39-amino-acid GLP-1 analog fused to an IgG4 Fc domain. Key features to confirm include the N-terminal pyroglutamate cyclization, the C-terminal amidation on the GLP-1 chain, and any disulfide or glycosylation sites in the Fc region. We must also consider that the fusion Fc (~50 kDa) introduces large polypeptide regions; our peptide sequencing must cover both the GLP-1 peptide and diagnostic fragments from the Fc. In summary, we focus on both the small GLP-1 segment (ensuring the 39-aa sequence is exact) and signature peptides from the Fc portion to confirm integrity of the whole API.
3.3 Enzymatic Digestion Strategy
To obtain fragments for sequencing, the dulaglutide API is enzymatically digested. Trypsin is the primary enzyme of choice, as it yields peptides suitable for MS/MS that cover the sequence. (For the Fc region, alternative proteases or specialized cleaving enzymes like IdeS can be used if needed, but trypsin alone often provides sufficient coverage for identity confirmation.) We perform reduction and alkylation to break disulfides first, then digest with trypsin under optimized conditions. The resulting peptide mixture is then cleaned up and injected into the LC–MS/MS system.
3.4 Experimental Procedure
We analyze the digest by high-resolution LC–MS/MS in data-dependent acquisition. In practice, the instrument is set to perform a full MS scan (e.g. m/z 300–2000) followed by MS/MS on the most abundant ions. Fragmentation (HCD or CID) generates b- and y-ions that map the peptide sequences. The MS/MS spectra are acquired with high resolution (e.g. <5 ppm mass accuracy) to distinguish any close mass variants. We include the RLD (reference) sample in parallel to ensure direct comparability. In summary, the workflow is: digest → LC separation → HRMS full-scan + MS/MS.
3.6 Data Analysis
Immediately after acquisition, we process the MS/MS data using specialized software (e.g. Byonic, PEAKS, UNIFI). The software matches each observed spectrum to the theoretical dulaglutide sequence, with known modifications (pyroglutamate, amidation, oxidation, deamidation, etc.) set as variable modifications. We require that every expected fragment ion be observed. For example, the N-terminal pyroglutamate yields a +111.03 Da shift vs. unmodified Glu, and this shift must appear correctly in the fragment series. The output is a peptide-spectrum map that covers the entire sequence. We manually review key fragments to ensure coverage is complete. Essentially, 100% coverage of the sequence by matched MS/MS fragments is required for sameness.
3.7 Key Observations / Acceptance Criteria
- Full sequence coverage: 100% of the dulaglutide amino-acid sequence is confirmed by MS/MS fragments (combined from all digests).
- No sequence variants: There are no extra or missing residues. Any unexpected fragment (not predicted by dulaglutide’s sequence) would indicate an error.
- Expected modifications confirmed: The N-terminal pyroglutamate and C-terminal amide must be correctly identified in the data (e.g. the +111 Da shift at the N-terminus).
- Mass accuracy: All parent ions and fragment ions fall within tight error (typically <5 ppm) of theoretical mass.
- Software scores: High confidence peptide-spectrum matches (low FDR).
Meeting these criteria proves the generic’s primary structure is identical to the RLD. Any discrepancy (e.g. missing fragment ions) triggers further investigation.
4. Peptide Mapping (Comparative Fingerprinting)
4.1 Objective
Peptide mapping (comparative fingerprinting) aims to compare the peptide fragment patterns of the generic and RLD digests in a single LC–MS assay. We create side-by-side maps of RLD vs generic to confirm that every tryptic peptide in the reference digest has an identical counterpart in the test sample. This LC–MS “fingerprint” simultaneously verifies sequence coverage and highlights any differences (missing peptides, extra peaks, co-eluting interferences).
4.2 Strategy
We digest the test and RLD samples in parallel under identical conditions. Each digest is injected on the same LC–MS system (typically a high-resolution Orbitrap/Q-TOF) using a reversed-phase gradient that separates the peptides. The LC method (e.g. C18 column at ~60°C, 0.1% formic acid buffers, gradient elution) is optimized for sharp, well-resolved tryptic peptides. During the LC run, the MS collects full-scan data (and optionally data-dependent MS/MS) across the gradient. The result is two chromatographic maps (RLD and test) of peptide masses vs retention time.
4.3 Experimental Workflow
- Sample Prep: Prepare parallel digests of the RLD and generic API using identical conditions. Standardize protein concentration so each run injects the same peptide amount.
- LC–MS Run: Inject RLD and test digests (separately) using the same instrument and method. For example, a typical setup might use a 2.1×100 mm, 1.7 µm C18 column at 60 °C, with a 5–40% acetonitrile gradient over 30 min. HRMS full scans are acquired (m/z ~300–2000) throughout. A reference peptide digest is run periodically for system suitability.
Example LC Conditions:
| Parameter | Setting |
|---|---|
| Column | BEH C18, 2.1×100 mm, 1.7 µm, 130 Å |
| Flow Rate | 0.3 mL/min |
| Temperature | 60 °C |
| Mobile Phase A | 0.1% formic acid in water |
| Mobile Phase B | 0.1% formic acid in acetonitrile |
| Gradient | 5–40% B over 30 min |
| Injection Volume | 2–5 µL (~2 µg peptide) |
4.4 Data Analysis
Data processing is automated using software (e.g. UNIFI or similar) with a library of expected peptide masses. The software overlays the RLD and test chromatograms (as two traces) and matches peaks by retention time and accurate mass. For each identified peptide in the RLD run, we check that the test run has a peak at the same retention time (within ±0.1 min) and m/z (within ppm tolerance). We also compare peak areas to ensure the relative abundances are similar. Key tasks:
- Peak Matching: Align each RLD peptide peak to a corresponding test peak by m/z and RT.
- Intensity Ratios: Check that the area of each matched peptide in the test sample is within ~20% of the RLD (large deviations may signal issues).
- New Peak Detection: Any peak present in the test that has no match in the RLD is flagged as a “new peak” (potential impurity or artifact). Such peaks are examined by MS/MS.
4.5 Key Observations / Acceptance Criteria
- Identical Chromatograms: All major peptides in the RLD map appear in the generic map with the same retention times and masses.
- No Unexplained Peaks: No significant new peaks in the generic that are absent in the RLD digest (beyond system noise). Any new peak is treated as an impurity.
- Consistent Ratios: Relative peak intensities match (within acceptable variance).
- System Suitability: Performance is confirmed by standard peptide digests.
Meeting these criteria confirms a successful fingerprinting comparison. If mismatches occur, it indicates a difference in composition or process impurity that must be addressed.
5. Impurity Profiling by HRMS
5.1 Objective
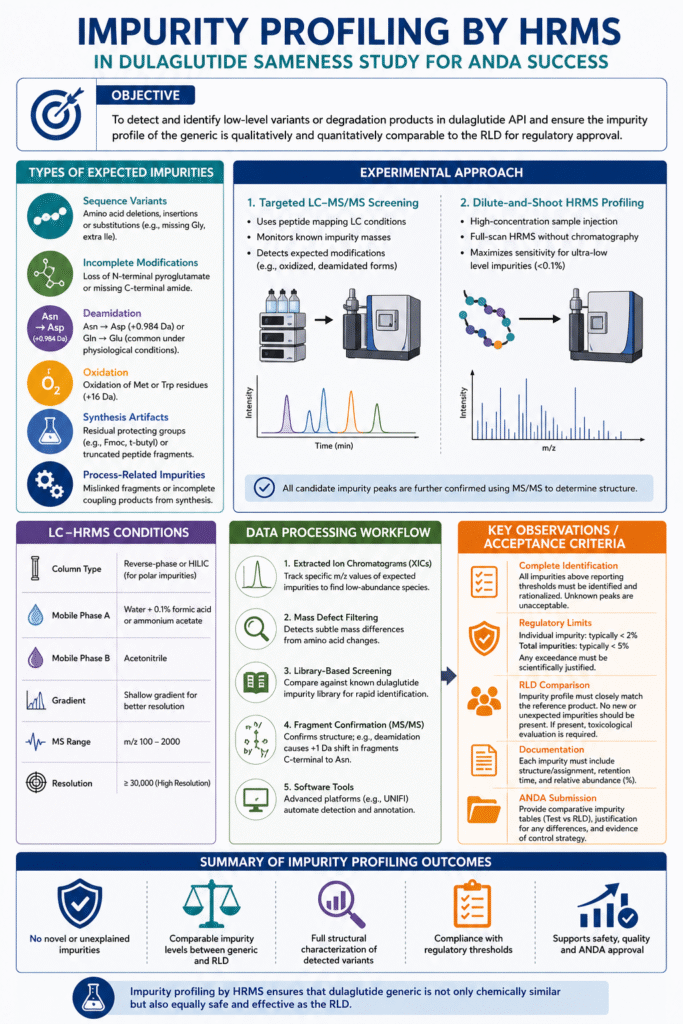
Impurity profiling aims to detect and identify any low-level variants or degradation products in the dulaglutide API. In other words, we scan beyond the main peptide to capture trace species that could affect safety. The FDA expects that the generic’s impurity profile be qualitatively and quantitatively comparable to the RLD. Using high-resolution MS (often coupled with LC), we look for truncated peptides, modified residues, synthetic by-products, or storage degradants.
5.2 Types of Expected Impurities
Based on peptide chemistry, common impurities include:
- Sequence variants: Single amino-acid deletions, insertions or substitutions (e.g. a missing Gly or extra Ile).
- Incomplete modifications: Loss of N-terminal pyroglutamate or missing C-terminal amide.
- Deamidation: Asn→Asp (mass +0.984 Da) or Gln→Glu (common at physiological pH).
- Oxidation: Met or Trp residues can oxidize (+16 Da).
- Synthesis artifacts: Residual protecting groups (e.g. Fmoc, t-butyl) or small peptide fragments from incomplete coupling.
- Process-related: Any truncation or mislinked fragments arising from the synthetic route.
We specifically check for these variants because even ppm-level differences can be critical for peptides.
5.3 Experimental Approach
We typically use two complementary HRMS methods for impurities:
- Targeted LC–MS/MS screening: Use the same LC separation as in peptide mapping, then mine the HRMS data for specific expected impurity masses. For example, we monitor masses corresponding to known modifications (deamidated or oxidized forms).
- Dilute-and-shoot HRMS profiling: Prepare a very high-concentration dulaglutide sample and run a full-scan HRMS (often without chromatography). This maximizes sensitivity to detect very low-level species (<0.1% of API).
In both cases, any candidate impurity peak is targeted for MS/MS to confirm its structure. High mass accuracy allows assignment of elemental composition (e.g. +0.9841 suggests deamidation) and the fragments verify the location of modification.
5.4 LC–HRMS Conditions
If using LC–MS, we may employ a shallow organic gradient. In some cases, a hydrophilic interaction (HILIC) column is used to better retain polar impurities (e.g. oxidized peptides). Typical mobile phases are 0.1% formic acid (or ammonium acetate) in water and acetonitrile. An HRMS instrument scans, for example, m/z 100–2000 at high resolution (≥30,000). This setup is sensitive enough to detect minor components that co-elute with the main peptide peak or elute separately.
5.5 Data Processing
Specialized workflows are used to extract potential impurity signals from the HRMS data. Key techniques include:
- Extracted Ion Chromatograms (XICs): We extract ions at the m/z of expected impurities (e.g. [M+H]+ of deamidated dulaglutide) to search chromatographically.
- Mass defect filtering: We scan the full-scan spectrum for peaks whose mass differences correspond to amino-acid changes.
- Library comparison: We compile a library of known dulaglutide variants (e.g. loss of pyroglutamate, deamidated sequences) and screen the data for matches.
- Fragment confirmation: For each candidate impurity, we trigger MS/MS (if needed) and check that fragment patterns match the hypothesized structure (e.g. N→D deamidation shifts all fragments C-terminal to Asn by +1 Da).
These methods, often implemented in software like UNIFI, help us pick out even co-eluting or low-abundance variants.
5.6 Key Observations / Acceptance Criteria
- Identification of all peaks: Every impurity signal above reporting thresholds must be identified and rationalized. Unidentified peaks indicate a gap in understanding.
- Regulatory limits: Individual impurities should generally be <2% and total impurities <5% (per pharmacopeia) unless justified. Any impurity exceeding limits must be explained or minimized.
- RLD comparison: The generic’s impurity profile should match the RLD’s. Any new impurity in the generic must be evaluated (and qualified toxicologically if needed). Ideally, impurity levels in the generic are ≤ those in the reference.
- Reporting: All impurities (name/structure and % area) are documented. For the ANDA, we provide a detailed comparison table and justify any minor differences.
In summary, a passing impurity profiling means no unexpected or novel modifications are present, and the impurity load is within acceptable bounds relative to the RLD.
6. Intact Mass Analysis
6.1 Objective
The objective of intact mass analysis is to verify the whole-molecule molecular weight of dulaglutide, ensuring it matches the theoretical mass and the reference drug. This is a rapid check on the complete peptide. Even a small mass difference (a few Daltons) can indicate an amino-acid change or post-translational difference. Thus, confirming the monoisotopic mass of the intact protein confirms “mass closure” and overall identity.
6.2 Method
We infuse or inject an undigested dulaglutide sample (no chromatography or a short trap column) into the HRMS. Dulaglutide (being ~60 kDa) will appear as multiple charged ions (e.g. +50 to +60 in ESI). We deconvolute the charge envelope to calculate the monoisotopic (zero-charge) mass. The observed mass is compared to the expected value (for dulaglutide, ≈59,669 Da from known sequence). The goal is to be within a tight tolerance (e.g. ±0.01%), typically well within modern HRMS accuracy.
6.3 Observations
We look for a single dominant peak corresponding to the intact protein. Sub-ppm accuracy is achievable, so a near-perfect match (±5 ppm) between measured and theoretical mass is expected. If the mass is exactly as expected, it confirms that the entire peptide (and Fc fusion) is present. Any deviation may point to a modification: for example, a +0.984 Da shift would suggest a deamidation, as noted above. In dulaglutide’s case, a precise intact mass match is a strong “initial confirmation” of identity.
7. NMR Characterization (Orthogonal Confirmation)
7.1 Objective
NMR spectroscopy is used as an orthogonal confirmation of molecular structure and purity. The objective is to verify the 3D chemical structure of dulaglutide in solution by examining its nuclear magnetic resonance signals. NMR serves as an independent check on the MS-based results: if the MS data match, the NMR should also overlay between generic and RLD, confirming the same peptide conformation and environment.
7.2 Techniques
We perform 1D and 2D NMR experiments on the peptide. This typically includes:
- 1D ^1H NMR: Gives the chemical-shift “fingerprint” of all proton types.
- 1D ^13C NMR: (if sensitivity allows) provides carbon backbone shifts.
- 2D COSY/TOCSY: Correlates protons through spin–spin coupling, helping identify spin systems of individual amino acids.
- 2D HSQC: Correlates ^1H to ^13C (or ^15N) and provides residue-specific information.
- NOESY/ROESY: (optional) reveals spatial proximities (used mainly for small peptides).
By combining these, we cover the aliphatic and aromatic regions, amide region, etc. In practice, due to dulaglutide’s size, we focus on regions like the GLP-1 segment and diagnostic Fc signals.
7.3 Key Focus Areas
Key NMR observables include chemical shifts and cross-peak patterns for signature residues. We check that the chemical shifts of diagnostic groups (e.g. alpha protons, aromatic side chains) match the reference. For example, the GLP-1 analog portion has a known helical motif whose backbone NHs and Cαs will appear in characteristic regions. The Fc region’s glycine and alanine peaks should also coincide. The absence of any extra peaks is critical (no new peaks means no additional compound). Consistent coupling patterns (COSY) ensure the same connectivity. All these checks verify that the generic’s higher-order structure is the same.
7.4 Acceptance Criteria
- Spectral overlap: The ¹H NMR spectrum of the generic overlays closely with that of the RLD (after minor chemical shift referencing). All major peaks coincide.
- No extra resonances: No additional signals are present in the test spectrum. Unexpected peaks would indicate an impurity or degradation product.
- Expected cross-peaks: 2D COSY/HSQC patterns match between test and reference for key residues.
- Chemical shift consistency: The chemical shifts (within ~0.1 ppm) align with reference values for known residues.
In line with guidelines, NMR serves to reinforce identity. Matching NMR spectra (1D/2D) provide orthogonal proof that the peptide’s structure and environment are unchanged.
8. Comparative Sameness Assessment (RLD vs Test)
After collecting all data, we perform a comprehensive side-by-side comparison of the generic and reference datasets. For every analytical measure above, the generic must meet the reference within acceptance criteria. In practice, this means overlaying spectra and chromatograms, compiling tables of peptide identities and impurities, and statistically confirming equivalence (where applicable).
All analytical datasets are compared side-by-side.
- This aligns with Sameness Evaluation in ANDA
- And Active Ingredient Sameness in ANDA
Successful studies show complete analytical equivalence.
For example, we overlay the LC–MS peptide maps (generic vs RLD) and highlight complete peak matches, or display the intact mass deconvolution with the target mass annotated. If all comparisons pass (100% sequence coverage, no new peaks, matching mass and spectra), we declare the samples “analytically indistinguishable.” This unified assessment – often summarized in an ANDA analytical report – conclusively demonstrates sameness. At that point, the generic is shown to be identical to the RLD in all relevant attributes, satisfying FDA/HC expectations.
9. Regulatory Considerations for ANDA
Regulatory agencies require a robust sameness package for peptide generics. In the US, synthetic peptide generics are submitted under 505(j) ANDAs (or 505(b)(2) if needed). Health Canada uses the ANDS pathway, but similarly mandates complete analytical proof of sameness. Both FDA and Health Canada demand confirmed amino-acid sequence, verified molecular mass, and a comparable impurity profile. In practice:
- FDA Guidance: FDA’s “Synthetic Peptide RLD” guidance (Nov 2021) advises how to determine if ANDA is appropriate. While dulaglutide is rDNA-derived, a fully synthetic generic with identical structure would generally be filed as an ANDA, provided all sameness criteria are met.
- Sequence Identity: As highlighted above, the FDA requires 100% sequence identity. Any sequence variants can lead to rejection.
- Impurity Matching: The generic’s impurity fingerprint must match the RLD’s, as qualitatively and quantitatively equivalent. Any novel impurity above threshold requires justification (often toxicology data).
- Orthogonal Confirmation: Both agencies expect orthogonal techniques (MS + NMR + others) to support equivalence.
- Bioequivalence: A traditional bioequivalence study is usually required for an ANDA, but it cannot substitute for analytical sameness. In fact, regulators typically insist on completing the analytical comparability first before considering BE .
- Reference Standards: The generic must be compared to a qualified reference standard (U.S. RLD or Canadian Reference Product) following jurisdictional rules.
Regulatory agencies require robust sameness data.
- FDA expectations are outlined in FDA Peptide Sameness Study Requirements
- Common pitfalls are discussed in Peptide Sameness Study Deficiencies
Key requirements:
- 100% sequence identity
- Comparable impurity profile
- Orthogonal confirmation
In summary, our analytical strategy is designed to fulfill these regulatory requirements. By rigorously documenting identity, purity, and structural equivalence, the generic dulaglutide is well-positioned for ANDA/ANDS approval without analytical deficiencies.
Conclusion:
A thorough Dulaglutide Sameness Study combines sequencing, mapping, HRMS/LC-MS profiling, intact-mass analysis, and NMR to definitively prove equivalence to the RLD. Each step—confirming 100% amino-acid sequence, matching peptide fingerprints, identifying trace impurities, verifying intact mass, and validating structure by NMR—contributes orthogonal evidence of sameness. When all acceptance criteria are met, regulators are assured that the generic and reference APIs are indistinguishable in quality and safety attributes. Such a scientifically rigorous approach not only aligns with FDA and Health Canada expectations, but also significantly reduces approval risk.
A Dulaglutide Sameness Study integrates sequencing, mapping, HRMS profiling, intact mass analysis, and NMR to prove equivalence.
- Comparable studies include Exenatide Sameness Study, Tirzepatide Sameness Study , and Retatrutide Sameness Study
This approach ensures regulatory compliance and minimizes ANDA approval risk.
ResolveMass Laboratories’ experienced scientists, FDA-registered facilities, and validated workflows provide the EEAT (Experience, Expertise, Authoritativeness, Trustworthiness) needed for a high-stakes ANDA submission. Our experts will ensure your dulaglutide generic is backed by comprehensive, regulator-ready data.
Frequently Asked Questions:
A Dulaglutide Sameness Study is an analytical comparison between a generic dulaglutide product and the Reference Listed Drug (RLD). It confirms structural, chemical, and purity equivalence required for ANDA approval. The study typically includes peptide sequencing, peptide mapping, impurity profiling, intact mass analysis, and NMR characterization. These methods collectively verify identity, quality, and consistency. It is a critical step for demonstrating pharmaceutical equivalence of peptide generics.
A Dulaglutide Sameness Study is important because the FDA requires generic manufacturers to prove equivalence to the RLD. Since dulaglutide is a complex peptide drug, simple assays are not sufficient. Detailed characterization helps establish sameness in sequence, impurities, and higher-order structure. This reduces regulatory risk and supports successful ANDA submission. Without strong analytical evidence, approval can be delayed or rejected.
A typical Dulaglutide Sameness Study uses multiple orthogonal techniques for complete characterization. These include HRMS/LC-MS for mass analysis, peptide mapping for fingerprinting, and MS/MS for sequence confirmation. Intact mass analysis verifies full molecular weight. 1D and 2D NMR are used for structural confirmation. Combining these methods strengthens confidence in comparability data.
Impurity profiling acceptance criteria generally require individual impurities to remain below specified thresholds, often below 2%. Total impurities are usually expected below 5%, unless otherwise justified. All significant impurities must be identified and characterized. The impurity profile should closely match the RLD. Any new impurity in the generic product requires additional evaluation.
Intact mass analysis measures the molecular weight of the full dulaglutide molecule without digestion. It confirms whether the generic product has the correct mass and expected modifications. This method can quickly identify truncations, adducts, or large modifications. It serves as a rapid orthogonal confirmation tool. Accurate intact mass matching is essential for sameness evaluation.
Reference
- Scott LJ. Dulaglutide: a review in type 2 diabetes. Drugs. 2020 Feb;80(2):197-208.https://link.springer.com/article/10.1007/s40265-020-01260-9
- Scheen AJ. Dulaglutide for the treatment of type 2 diabetes. Expert opinion on biological therapy. 2017 Apr 3;17(4):485-96.https://www.tandfonline.com/doi/abs/10.1080/14712598.2017.1296131
- Gerstein HC, Colhoun HM, Dagenais GR, Diaz R, Lakshmanan M, Pais P, Probstfield J, Riesmeyer JS, Riddle MC, Rydén L, Xavier D. Dulaglutide and cardiovascular outcomes in type 2 diabetes (REWIND): a double-blind, randomised placebo-controlled trial. The Lancet. 2019 Jul 13;394(10193):121-30.https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(19)31149-3/abstract
- Sanford M. Dulaglutide: first global approval. Drugs. 2014 Nov;74(17):2097-103.https://link.springer.com/article/10.1007/s40265-014-0320-7


