Introduction:
GLP-1 Enzymatic Digestion Mapping is a structured analytical process where enzymes are used to cleave GLP-1 peptides into smaller fragments for detailed structural analysis.
At ResolveMass Laboratories Inc., this approach is a critical part of advanced analytical characterization of GLP-1 peptide drugs, enabling high-confidence sequence verification, impurity profiling, and regulatory-grade reporting.
With the growing importance of GLP-1 peptides in therapeutics (especially metabolic and diabetes treatments), precise enzymatic digestion strategies are essential for ensuring accuracy, reproducibility, and compliance. These workflows are closely aligned with regulatory requirements for GLP-1 peptide characterization.
Share via:
Summary:
- GLP-1 Enzymatic Digestion Mapping is essential for accurate structural characterization of GLP-1 peptides.
- Enzymatic digestion breaks peptides into predictable fragments for mass spectrometry analysis.
- Key enzymes include Trypsin, Glu-C, Chymotrypsin, and Asp-N, each offering unique cleavage specificity.
- A multi-enzyme strategy improves sequence coverage and confidence, similar to advanced GLP-1 analog peptide sequencing workflow approaches.
- Optimized workflows ensure high reproducibility, regulatory compliance, and minimal analytical errors.
- Expert validation and orthogonal approaches significantly enhance data reliability, as seen in de novo GLP-1 peptide sequencing accuracy methodologies.
1: Why is Enzymatic Digestion Important in GLP-1 Peptide Mapping?
Enzymatic digestion is crucial because it breaks down complex GLP-1 peptides into smaller, well-defined fragments that can be accurately analyzed using mass spectrometry, forming a key step in LC-MS characterization of GLP-1 peptides.
Key Benefits:
- Enables precise sequence identification, supporting peptide sequencing of GLP-1 peptide workflows
- Improves mass spectrometry resolution by reducing spectral complexity
- Helps detect modifications and impurities, critical for GLP-1 peptide impurity characterization
- Ensures complete structural characterization for regulatory and research purposes
Without proper enzymatic digestion, peptide samples produce overlapping signals and highly complex spectra, making it difficult to achieve accurate identification and reliable analytical results.
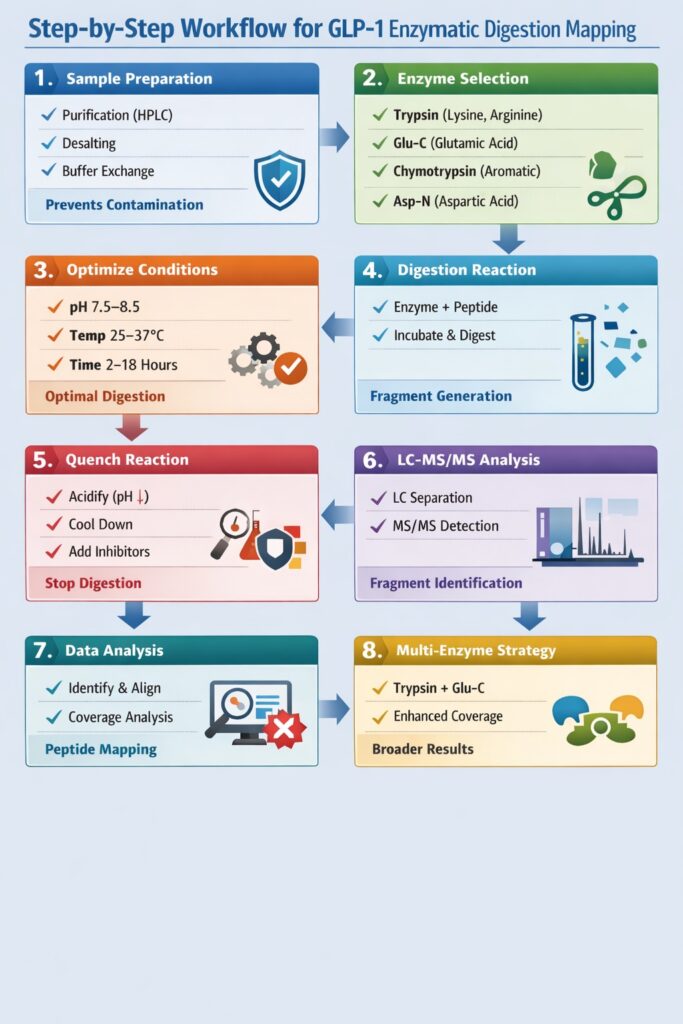
2: Step-by-Step Workflow for GLP-1 Enzymatic Digestion Mapping
Step 1: Sample Preparation
Proper sample preparation ensures peptide purity and optimal digestion efficiency.
Before enzymatic digestion, GLP-1 samples must be carefully prepared to eliminate contaminants and stabilize the peptide structure.
Key Steps:
- Peptide purification (HPLC or equivalent)
- Desalting to remove interfering salts
- Buffer exchange to maintain enzyme compatibility
Why It Matters:
- Prevents enzyme inhibition
- Enhances digestion efficiency
- Improves mass spectrometry signal quality
This step is foundational for accurate peptide sequencing of GLP-1 drugs.
Step 2: Selection of Appropriate Enzymes
Choosing the right enzyme determines cleavage specificity and overall mapping accuracy.
Different enzymes cleave at specific amino acid residues, producing predictable and analyzable peptide fragments.
Common Enzymes Used:
| Enzyme | Cleavage Specificity | Application |
|---|---|---|
| Trypsin | Lysine (K), Arginine (R) | Standard mapping |
| Glu-C | Glutamic acid (E) | Complementary mapping |
| Chymotrypsin | Aromatic residues (F, Y, W) | Hydrophobic regions |
| Asp-N | N-terminal of Aspartic acid (D) | Alternative cleavage |
Best Practice:
- Use multiple enzymes to improve sequence coverage and reduce ambiguity.
Use multiple enzymes to improve sequence coverage and reduce ambiguity—especially important when addressing GLP-1 peptide sequencing challenges.
Step 3: Optimization of Digestion Conditions
Controlled conditions ensure efficient and reproducible enzymatic cleavage.
Each enzyme requires specific environmental conditions for optimal activity.
Key Parameters:
- pH: Typically 7.5–8.5 (varies by enzyme)
- Temperature: 25–37°C
- Incubation Time: 2–18 hours
- Enzyme-to-Substrate Ratio: Optimized per protocol
Why Optimization Matters:
- Prevents over-digestion or incomplete digestion
- Ensures consistent fragment generation
- Reduces experimental variability
Optimization plays a key role in achieving reliable results in GLP-1 peptide sequencing analytical technique workflows.
Step 4: Enzymatic Digestion Reaction
This step involves controlled cleavage of GLP-1 peptides into defined fragments.
This step generates predictable peptide fragments for downstream analysis and supports high-confidence GLP-1 peptide sequencing CRO services.
The enzyme is added to the prepared sample under optimized conditions.
Process Overview:
- Mix enzyme with peptide sample
- Incubate under controlled conditions
- Monitor digestion progress (optional)
Output:
- A mixture of predictable peptide fragments ready for analysis
Step 5: Quenching the Reaction
Digestion is stopped to preserve fragment integrity.
Once digestion is complete, the reaction must be terminated to prevent further cleavage.
Maintains fragment integrity and prevents over-digestion artifacts, which is essential for downstream GLP-1 peptide impurity sequencing analysis.
Common Methods:
- Lowering pH (acidification)
- Cooling the sample
- Adding inhibitors
Importance:
- Maintains fragment stability
- Prevents over-digestion artifacts
Step 6: LC-MS/MS Analysis
Fragmented peptides are analyzed using high-resolution mass spectrometry.
Liquid Chromatography–Mass Spectrometry (LC-MS/MS) separates and identifies peptide fragments.
This aligns with advanced LC-MS characterization of GLP-1 peptides for accurate detection and identification.
Key Components:
- LC Separation: Reduces sample complexity
- MS Detection: Measures mass-to-charge ratios
- MS/MS Fragmentation: Provides sequence information
Outcome:
- High-resolution data for peptide identification and mapping
Step 7: Data Analysis and Peptide Mapping
Software and expert review reconstruct the peptide sequence.
Advanced algorithms interpret MS/MS data to identify peptide fragments and map them back to the original GLP-1 sequence.
Often integrated with peptide sequencing of GLP-1 peptide workflows for full structural validation.
Key Steps:
- Fragment identification
- Sequence alignment
- Coverage analysis
Best Practice:
- Combine automated analysis with expert validation for maximum accuracy
Step 8: Multi-Enzyme Strategy for Enhanced Coverage
Using multiple enzymes increases sequence coverage and confidence.
A single enzyme may not provide complete coverage of the GLP-1 peptide.
Multi-enzyme strategies improve reliability and reduce ambiguity, a best practice commonly used by providers offering CRO for GLP-1 peptide characterization.
Advantages:
- Covers hard-to-digest regions
- Confirms ambiguous sequences
- Improves overall mapping reliability
Example Strategy:
- Trypsin + Glu-C combination
- Sequential or parallel digestion workflows
3: Best Practices for GLP-1 Enzymatic Digestion Mapping
Following best practices ensures high accuracy and compliance with GLP-1 peptide characterization regulatory requirements.
Additionally:
- Stability considerations should align with GLP-1 peptide stability analytical methods
- Outsourcing complex workflows can benefit from outsource GLP-1 peptide sequencing services
To achieve high-quality and reliable results, implement the following expert recommendations:
1. Use High-Purity Enzymes
High-purity enzymes minimize unwanted side reactions and improve digestion specificity.
- Use analytical-grade or sequencing-grade enzymes
- Avoid contaminants that may interfere with cleavage
- Store enzymes under recommended conditions to maintain activity
2. Validate Digestion Efficiency
Validation confirms that digestion is complete and reproducible.
- Perform pilot or test runs before full analysis
- Monitor digestion completeness using LC-MS
- Check for missed cleavages or unexpected fragments
3. Optimize Protocols for Each Peptide
Each GLP-1 peptide may require customized digestion conditions.
- Adjust pH, temperature, and incubation time as needed
- Optimize enzyme-to-substrate ratios
- Avoid assuming a “one-size-fits-all” workflow
4. Implement Orthogonal Techniques
Using complementary analytical methods improves confidence in results.
- Combine LC-MS/MS with additional techniques (e.g., different fragmentation modes)
- Cross-validate peptide identification and modifications
- Reduce analytical bias and increase data robustness
5. Include Expert Review
Expert validation ensures data accuracy and regulatory readiness.
- Perform manual review of complex or ambiguous spectra
- Confirm sequence assignments and modifications
- Ensure results meet regulatory and quality standards
By applying these best practices, GLP-1 Enzymatic Digestion Mapping becomes more reliable, reproducible, and suitable for high-confidence peptide characterization.
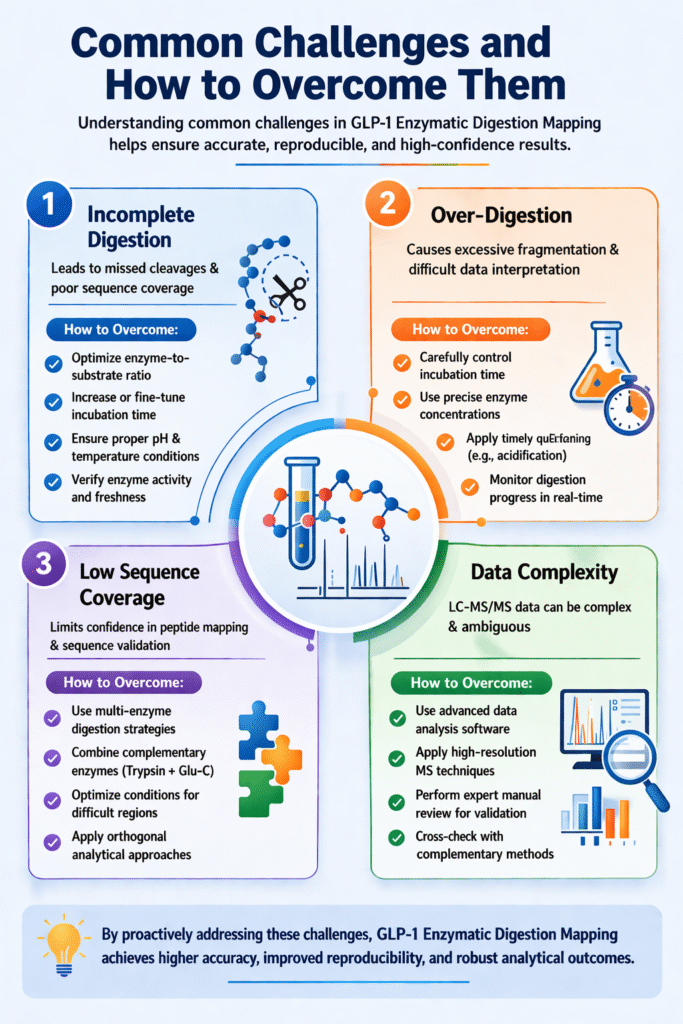
4: Common Challenges and How to Overcome Them
Understanding common challenges in GLP-1 Enzymatic Digestion Mapping helps ensure accurate, reproducible, and high-confidence results.
Challenges such as incomplete digestion, over-digestion, and data complexity are commonly encountered in GLP-1 peptide sequencing challenges.
Addressing these requires:
- Advanced analytical tools
- Multi-enzyme strategies
- Expert validation
- Integration with impurity-focused workflows like GLP-1 peptide impurity characterization
1. Incomplete Digestion
Incomplete digestion leads to missed cleavages and poor sequence coverage.
How to Overcome:
- Optimize enzyme-to-substrate ratio
- Increase or fine-tune incubation time
- Ensure proper pH and temperature conditions
- Verify enzyme activity and freshness
2. Over-Digestion
Over-digestion causes excessive fragmentation, making data interpretation difficult.
How to Overcome:
- Carefully control incubation time
- Use precise enzyme concentrations
- Apply timely quenching methods (e.g., acidification)
- Monitor digestion progress during the reaction
3. Low Sequence Coverage
Low coverage limits confidence in peptide mapping and sequence validation.
How to Overcome:
- Use multi-enzyme digestion strategies
- Combine complementary enzymes (e.g., Trypsin + Glu-C)
- Optimize digestion conditions for difficult regions
- Apply orthogonal analytical approaches
4. Data Complexity
Complex LC-MS/MS data can lead to ambiguous or incorrect interpretations.
How to Overcome:
- Use advanced data analysis software and algorithms
- Apply high-resolution mass spectrometry techniques
- Perform manual expert review for validation
- Cross-check results using complementary methods
By proactively addressing these challenges, GLP-1 Enzymatic Digestion Mapping can achieve higher accuracy, improved reproducibility, and robust analytical outcomes.
5: How ResolveMass Laboratories Inc. Ensures High-Quality Results
ResolveMass Laboratories Inc. ensures high-quality results by combining advanced instrumentation, optimized workflows, and deep expertise in analytical characterization of GLP-1 peptide drugs.
At ResolveMass Laboratories Inc., enzymatic digestion workflows are developed with precision, consistency, and regulatory alignment to deliver reliable analytical outcomes for complex peptide characterization.
Their capabilities include:
- Advanced LC-MS/MS platforms
- Customized digestion strategies
- Expertise in sequencing and mapping
- Strong compliance with regulatory requirements for GLP-1 peptide characterization
Key Strengths
1. Advanced High-Resolution Mass Spectrometry Platforms
- Utilizes cutting-edge LC-MS/MS systems for accurate mass detection
- Enables high sensitivity and resolution for complex peptide mixtures
- Supports confident identification of sequence variants and modifications
2. Customized Multi-Enzyme Digestion Strategies
- Tailored enzyme selection based on peptide structure
- Improves sequence coverage and reduces analytical ambiguity
- Ensures comprehensive GLP-1 Enzymatic Digestion Mapping
3. Deep Expertise in Peptide Mapping and Sequencing
- Experienced scientists with strong domain knowledge
- Proven workflows for complex peptide characterization
- Expertise in both automated and manual data interpretation
4. Strong Focus on Data Validation and Compliance
- Multi-level data verification for accuracy and reproducibility
- Alignment with regulatory expectations and industry standards
- Detailed, audit-ready reporting for clients
They are also a trusted partner for GLP-1 peptide sequencing CRO services.
This integrated approach ensures clients receive accurate, reproducible, and regulatory-ready analytical data, making ResolveMass Laboratories Inc. a trusted partner for GLP-1 peptide mapping and advanced analytical services.
Conclusion:
GLP-1 Enzymatic Digestion Mapping is a foundational step in peptide characterization, enabling accurate sequence identification and structural validation.
By integrating optimized workflows, advanced peptide sequencing of GLP-1 peptide strategies, and regulatory-aligned methodologies, researchers can achieve high-confidence results.
As demand for GLP-1 therapeutics continues to rise, adopting robust analytical approaches—including peptide mapping GLP-1 peptides—is essential for ensuring quality, safety, and compliance.
Frequently Asked Questions:
GLP-1 Enzymatic Digestion Mapping is a structured analytical technique used to break down GLP-1 peptides into smaller, well-defined fragments using specific enzymes. These fragments are then analyzed using advanced techniques like LC-MS/MS to reconstruct the peptide structure. This approach enables accurate sequence verification and structural characterization. It is widely used in pharmaceutical research, especially for GLP-1-based therapeutics. The method also supports impurity detection and modification analysis. Overall, it plays a critical role in ensuring data accuracy and reliability.
Enzymatic digestion is essential because GLP-1 peptides are relatively complex molecules that cannot be easily analyzed in their intact form. By breaking them into smaller fragments, it becomes easier to identify and study their structure. This improves mass spectrometry resolution and reduces signal overlap. It also helps detect impurities, sequence variations, and post-translational modifications. Without digestion, analytical results may be inaccurate or incomplete. Therefore, it is a foundational step in peptide characterization workflows.
Several enzymes are commonly used in GLP-1 digestion mapping, each with specific cleavage properties. Trypsin is widely used as it cleaves at lysine and arginine residues. Glu-C targets glutamic acid residues, providing complementary fragmentation. Chymotrypsin is useful for hydrophobic and aromatic amino acids. Asp-N cleaves at the N-terminal side of aspartic acid residues. Using multiple enzymes ensures better sequence coverage and reduces analytical gaps. This combination improves confidence in peptide mapping results.
LC-MS/MS is a critical analytical technique used after enzymatic digestion. Liquid chromatography (LC) separates peptide fragments based on their chemical properties. Mass spectrometry (MS) then detects and measures the mass-to-charge ratio of these fragments. MS/MS fragmentation provides detailed structural information about each peptide. This allows researchers to reconstruct the original GLP-1 sequence accurately. It also helps identify impurities and modifications. Overall, LC-MS/MS ensures high-resolution and reliable analytical data.
There are several challenges in GLP-1 enzymatic digestion mapping that can affect accuracy. Incomplete digestion can lead to missed cleavages and poor sequence coverage. Over-digestion may cause excessive fragmentation, making analysis difficult. Data complexity from LC-MS/MS can also result in ambiguous interpretations. Additionally, some peptide regions may be difficult to digest due to structural constraints. These challenges require careful optimization and expert validation. Addressing them is essential for reliable results.
Digestion efficiency can be significantly improved by optimizing experimental conditions. Key factors include pH, temperature, and incubation time, which must match enzyme requirements. The enzyme-to-substrate ratio should also be carefully controlled. Using high-purity, sequencing-grade enzymes reduces unwanted side reactions. Proper sample preparation, including desalting and purification, is equally important. Monitoring digestion progress using LC-MS helps ensure completeness. These steps collectively enhance reproducibility and accuracy.
A multi-enzyme strategy is used to overcome limitations of single-enzyme digestion. Each enzyme cleaves at specific amino acid residues, so using multiple enzymes generates complementary fragments. This improves sequence coverage and helps analyze difficult regions. It also reduces ambiguity in peptide identification. Multi-enzyme approaches provide cross-validation of results, increasing confidence. They are especially useful for complex GLP-1 analogs. Overall, this strategy enhances mapping reliability.
Reference
- Ludwig MQ, Todorov PV, Egerod KL, Olson DP, Pers TH. Single-cell mapping of GLP-1 and GIP receptor expression in the dorsal vagal complex. Diabetes. 2021 Sep 1;70(9):1945-55.https://diabetesjournals.org/diabetes/article-abstract/70/9/1945/137747
- Manjunath PG, Ravi NK, Sridhar SK, Raghavendra NN. Innovative strategies and nanocarrier approaches for enhancing the oral bioavailability of macromolecular therapeutics. Iranian Journal of Basic Medical Sciences. 2025;28(12):1607.https://pmc.ncbi.nlm.nih.gov/articles/PMC12829707/
- Xu Y, Huang T, Zhang Y, Ji D, Tuttle KR, Carrero JJ, Fu EL. Risk of heart failure hospitalization for GLP-1 receptor agonists versus DPP-4 inhibitors or SGLT-2 inhibitors in patients with type 2 diabetes: a target trial emulation. Circulation. 2026 Feb 24.https://www.ahajournals.org/doi/abs/10.1161/CIRCULATIONAHA.125.075157
- Zhang S, Li ZM, Feng Y, Yu S, Li Z, Zhang D, Wang C. Dpp-iv inhibitory peptides from coix seed prolamins: Release, identification, and analysis of the interaction between key residues and enzyme domains. Journal of Agricultural and Food Chemistry. 2023 Sep 25;71(40):14575-92.https://pubs.acs.org/doi/abs/10.1021/acs.jafc.3c02733


