Introduction:
GLP-1 Peptide Mapping Using LC-MS is a powerful analytical approach used to confirm sequence integrity, identify modifications, and detect impurities in GLP-1 analogs. This method combines multi-protease digestion with high-resolution tandem mass spectrometry to achieve full sequence coverage and precise localization of acylation sites. The strategy uses accurate mass measurements from Orbitrap or Q-TOF systems to verify both amino acid sequences and synthetic modifications such as fatty acid side chains. It is especially useful for lipidated peptides, where structural complexity makes traditional mapping difficult. In addition, GLP-1 Peptide Mapping Using LC-MS allows simultaneous confirmation of molecular integrity, modification stability, and impurity detection in a single run.
Explore our specialized services: Peptide Characterization Techniques and Applications
Share via:
Technical Foundations of GLP-1 Peptide Mapping Using LC-MS
The workflow for GLP-1 Peptide Mapping Using LC-MS depends on controlled fragmentation of the peptide backbone while keeping side-chain modifications intact during ionization and chromatographic separation. GLP-1 analogs behave differently from standard peptides because they contain both hydrophilic and hydrophobic regions. The peptide backbone is water-soluble, while the fatty acid chain is highly hydrophobic, creating challenges during separation and ionization. This mixed behavior can lead to aggregation, surface adsorption, and uneven signal intensity across fragments. To overcome this, analysts optimize buffer composition, temperature, and organic solvent content. Careful control of these conditions improves solubility, signal stability, and reproducible fragmentation patterns.
Structural Engineering and Analytical Implications
Modern GLP-1 analogs are designed with chemical modifications that extend half-life and improve stability. Liraglutide, for instance, shares high sequence similarity with native GLP-1 but includes a palmitic acid attached to Lys26 through a glutamic acid spacer. Semaglutide contains additional changes such as an Aib substitution at position 8 and a C18 diacid chain, increasing hydrophobicity and resistance to enzymatic degradation. These structural changes influence chromatographic retention, digestion efficiency, and fragmentation behavior. Because of this, GLP-1 Peptide Mapping Using LC-MS must be customized for each analog to accurately identify backbone and modification sites. Precise mapping also supports batch comparability and confirms structural integrity during manufacturing.
| GLP-1 Analog | Molecular Weight (Da) | Key Modifications | Analytical Challenge |
|---|---|---|---|
| Liraglutide | 3751.2 | Lys26 palmitoylation (C16) | High propensity for surface adsorption. |
| Semaglutide | 4113.6 | Lys26 C18 diacid; Aib8 substitution | Extreme hydrophobicity; protease resistance at Lys26. |
| Tirzepatide | 4813.5 | Dual acylation (C20); Aib substitutions | Large size; complex fragmentation patterns. |
| Exenatide | 4186.6 | 39 amino acids; C-terminal amidation | C-terminal sequence verification. |
Deep dive into specific analog mapping: Peptide Mapping Study of Semaglutide
Optimization of Sample Preparation for Hydrophobic Peptides
Effective sample preparation for GLP-1 Peptide Mapping Using LC-MS requires strong chaotropic agents and balanced organic-aqueous solvent systems. Acylated peptides like semaglutide can form aggregates or stable secondary structures in aqueous buffers, which reduces digestion efficiency. Adding 6M urea or 6M guanidine-HCl disrupts these interactions and exposes cleavage sites for enzymes. Proper solubilization improves reproducibility and minimizes sample loss due to adsorption. Controlled denaturation also helps maintain modification stability while preparing peptides for digestion.
Denaturation, Reduction, and Alkylation Kinetics
Denaturation conditions must be optimized to avoid unwanted reactions such as carbamylation from overheated urea solutions. For GLP-1 analogs containing disulfide bonds, reduction with 10 mM DTT followed by alkylation with iodoacetamide linearizes the peptide. Alkylation should be performed in low-light conditions to prevent side reactions. Complete reduction and alkylation stabilize cysteine residues and improve chromatographic peak shapes. These steps are essential for reliable downstream LC-MS analysis.
Peptide-S-S-Peptide + 2R-SH → 2Peptide-SH + R-S-S-R
Peptide-SH + ICH2CONH2 → Peptide-S-CH2CONH2 + HI
These reactions must reach >99% completion to avoid partially reduced or alkylated species. Incomplete reactions can create overlapping signals and complicate impurity analysis. Full conversion improves chromatographic clarity and reduces ambiguity. This also increases confidence when detecting low-level degradants. Reliable preparation directly supports high-quality LC-MS results.
Learn more about our core capabilities: Peptide Characterization Service in Canada
Multi-Enzyme Digestion Strategies for 100% Sequence Coverage
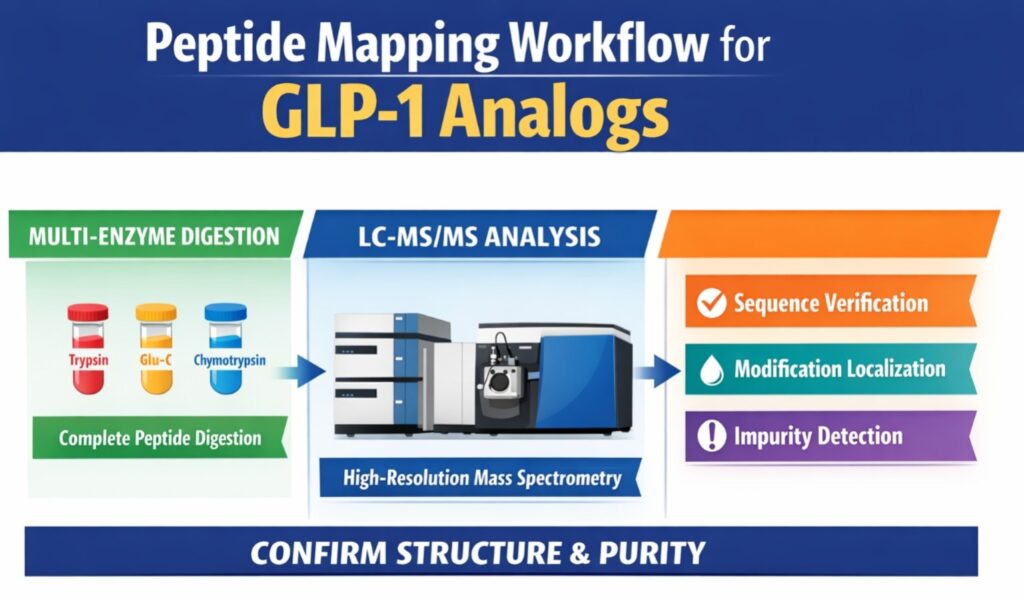
Complete sequence coverage in GLP-1 Peptide Mapping Using LC-MS is typically achieved using parallel digestion with Trypsin, Glu-C, and Chymotrypsin. Acylation at lysine residues often blocks tryptic cleavage, producing large fragments that are difficult to analyze. Using multiple enzymes generates overlapping fragments that cover these blocked regions. This improves redundancy and increases confidence in modification localization. Multi-enzyme digestion also enhances detection of low-abundance impurities.
Compare analytical methodologies: Peptide Mapping vs. Peptide Sequencing: Key Differences
Selection and Rationale for Alternative Proteases
Glu-C cleaves at glutamic acid and aspartic acid residues that are usually unaffected by acylation. Chymotrypsin targets hydrophobic residues, making it useful for lipidated regions. Asp-N helps with N-terminal characterization and stability mapping. Together, these enzymes produce overlapping fragments that enable complete sequence confirmation. This strategy reduces blind spots near modified residues.
| Enzyme | Optimal pH | Preferred Cleavage Site | Role in GLP-1 Mapping |
|---|---|---|---|
| Trypsin | 7.5 – 8.5 | Arg (R), Lys (K) | Efficient backbone cleavage; specific for non-modified regions. |
| Glu-C | 7.8 | Glu (E), Asp (D) | Overcomes K-acylation blocks; provides mapping for acidic regions. |
| Chymotrypsin | 8.0 | Phe (F), Trp (W), Tyr (Y) | Maps hydrophobic cores; essential for coverage of lipidated segments. |
| Asp-N | 6.0 – 8.5 | N-terminal to Asp (D) | Useful for specific N-terminal variants and stability mapping. |
Immobilized enzyme systems such as SMART Digest can reduce digestion time to less than one hour. These systems minimize autolysis and improve reproducibility. Faster digestion supports high-throughput workflows in quality control laboratories. Reduced variability improves comparability across batches.
Advanced Chromatographic Separation of Modified Fragments
Chromatographic separation in GLP-1 Peptide Mapping Using LC-MS is typically performed using wide-pore C4 or C18 columns operated at elevated temperatures. Running columns at 60°C to 80°C reduces viscosity and improves mass transfer for bulky lipidated fragments. This produces narrower peaks and better separation of closely related species. High-temperature operation also reduces non-specific adsorption and improves recovery. These benefits are particularly important for hydrophobic GLP-1 analogs.
Understand the importance of purity: Peptide Purity Testing Services
Column Chemistry: C4 vs. C18 for Hydrophobic Peptides
Traditional C18 columns often retain hydrophobic peptides too strongly, leading to carry-over between injections. C4 columns provide weaker retention and improved elution of lipidated fragments. This reduces carry-over while maintaining sequence coverage. Lower adsorption also improves reproducibility between runs. For many acylated GLP-1 analogs, C4 chemistry offers a more reliable solution.
Mobile Phase Modifiers and Gradient Optimization
Acidic modifiers control peptide protonation and improve peak shape. TFA provides strong separation but can suppress MS signal. DFA offers similar resolution with better sensitivity. Shallow gradients help resolve deamidated and isomeric impurities. Balanced solvent composition maintains solubility and stable ionization.
| Parameter | Optimized Setting | Technical Justification |
| Stationary Phase | C4 RP (sub-2 \mum) | Minimizes carry-over of acylated fragments. |
| Column Temp | 60°C – 70°C | Improves mass transfer and peak symmetry. |
| Modifier | 0.1% DFA or 0.1% FA | Balances ion-pairing with MS sensitivity. |
| Gradient Slope | 0.5% – 1.0% B/min | Necessary for resolving deamidated/isomeric impurities. |
| Organic Solvent | Acetonitrile/Methanol (50/50) | Enhanced solubility for lipidated species. |
Mass Spectrometry Parameters and Fragmentation Mechanics
Mass spectrometric detection in GLP-1 Peptide Mapping Using LC-MS uses high-resolution analyzers operating at ≥60,000 FWHM. High mass accuracy allows differentiation between fragments with very similar composition. This is important for identifying oxidation (+15.9949 Da) and deamidation (+0.9840 Da). Accurate isotopic envelopes improve confidence in assignments. These settings support reliable detection of low-level impurities.
Access high-resolution sequencing data: Semaglutide Peptide Sequencing via LC-MS/MS
Higher-Energy C-Trap Dissociation (HCD) and Electron-Transfer Dissociation (ETD)
HCD generates strong b- and y-ion series for backbone sequencing. However, lipidated side chains can reduce coverage. ETD complements HCD by preserving labile modifications and cleaving different bonds. Combining both methods improves sequence coverage and modification localization. This hybrid strategy is useful for complex GLP-1 analogs.
| MS Setting | Recommended Value | Impact on Data Quality |
|---|---|---|
| MS1 Resolution | 60,000 – 120,000 | Definitive identification of the precursor ion. |
| MS2 Resolution | 15,000 – 30,000 | Fast scan rates to maximize fragment-ion capture. |
| Collision Energy | 25 – 35 (NCE) | Optimized for b/y ion series without excessive internal fragmentation. |
| AGC Target | 1 × 10^6 – 3 × 10^6 | Optimizes the ion population for dynamic range and mass accuracy. |
| Isolation Window | 1.3 – 2.0 m/z | Minimizes chimeric spectra while maintaining sensitivity. |
Dynamic exclusion prevents repeated fragmentation of abundant peptides and increases detection of minor variants. This improves impurity profiling and sequence completeness. Optimized acquisition parameters enhance reproducibility. These controls are essential for regulated workflows.
Site-Specific Localization of Acylation and Spacers
Acylation sites are localized by identifying diagnostic mass shifts and confirming flanking fragment ions. For semaglutide, Lys26 includes gamma-glutamic acid and AEEA spacers linked to a C18 diacid. Detection of spacer-specific fragments confirms structural integrity. This ensures correct placement of lipid modifications. Accurate localization is critical for biosimilar comparability.
Technical workflow overview: GLP-1 Analog Peptide Sequencing Workflow
Detecting Isotope Distribution and Mass Errors
Isotope distribution analysis validates fragment identity and reduces isobaric interference. Observed patterns must match theoretical distributions with high fidelity. Mass errors are typically maintained below 3 ppm for Orbitrap systems. Continuous monitoring improves confidence in structural assignments. These checks support regulatory documentation.
Mapping Impurities and Degradation Products in GLP-1 Analogs
Impurity characterization in GLP-1 Peptide Mapping Using LC-MS relies on detecting mass shifts and retention changes. Common degradants include deamidation, oxidation, and truncation. Even small amounts of degradation can affect safety and efficacy. Accurate mapping enables early detection and control. This supports long-term stability studies.
The Challenge of D-Isomers and “n-1” Truncations
D-isomers and “n-1” truncations are difficult to separate because they have similar mass and hydrophobicity. Advanced UHPLC strategies with shallow gradients improve separation. Core-shell particles increase peak capacity and resolution. High-resolution MS confirms identity after chromatographic separation. Combined approaches improve impurity detection.
| Impurity Type | Mass Shift (Da) | Common Residues | Analytical Detection Strategy |
|---|---|---|---|
| Deamidation | +0.9840 | Asn (N), Gln (Q) | Shift in retention time; diagnostic b/y fragments. |
| Oxidation | +15.9949 | Met (M), Trp (W) | +16 Da mass shift; earlier elution (usually). |
| Truncation | Variable | N- or C-terminal | Targeted mass search; MS/MS sequence verification. |
| Isomerization | +0.0000 | Asp (D) | Retention time shift; unique ETD fragments. |
Forced degradation studies validate detection capability down to 0.1%. These studies simulate oxidation and deamidation conditions. Verified methods improve regulatory acceptance. Robust validation ensures analytical reliability.
Regulatory Validation and Sameness Studies (ICH Q6B)
Regulatory validation for GLP-1 Peptide Mapping Using LC-MS includes comparability studies demonstrating structural identity with the reference product. ICH Q6B requires full sequence confirmation and impurity characterization. LC-MS peptide mapping provides direct structural evidence. These studies support biosimilar and generic development. Comprehensive validation ensures compliance.
Meet regulatory standards: FDA Requirements for Peptide Characterization
ANDA Sameness Criteria for Generic Peptides
FDA guidance requires demonstration of API sameness between generic peptides and the reference drug. This includes identical primary sequence and comparable higher-order structure. Impurities above 0.5% must be justified. Detailed LC-MS mapping supports these requirements. Consistent analytical performance is essential for approval.
| Validation Attribute | Regulatory Requirement | Methodological Verification |
| Sequence Identity | 100% Confirmation | Overlapping peptide maps (LC-MS/MS). |
| Mass Accuracy | \le 10 ppm Error | HRAM-MS (Orbitrap/Q-TOF). |
| Purity | \ge 98\% typically | Chromatographic purity (LC-UV/MS). |
| Batch Consistency | 3 Batches Minimum | Replicate analysis across multiple drug substance batches. |
Troubleshooting Common Failures in GLP-1 Peptide Mapping
Reliable GLP-1 Peptide Mapping Using LC-MS requires monitoring system suitability metrics. Retention drift, adsorption, and ion suppression can affect results. Environmental factors strongly influence hydrophobic peptides. Routine checks help maintain consistency. Early troubleshooting prevents analytical failures.
Prepare for clinical filing: Peptide Characterization for IND and NDA
Managing Adsorption and Recovery Issues
Hydrophobic GLP-1 analogs often adsorb to plastic and metal surfaces. This leads to inconsistent recovery and carry-over. Using low-bind consumables reduces adsorption. Adding organic solvent or mild surfactants improves recovery. These adjustments enhance reproducibility.
Addressing Low Sequence Coverage in Acylated Regions
Low coverage near acylation sites often results from incomplete digestion. Increasing enzyme-to-substrate ratio improves cleavage. Adding acetonitrile enhances solubility of lipidated fragments. Sequential digestion further breaks down large peptides. These strategies improve MS detectability.
Future Outlook: Automation and the Multi-Attribute Method (MAM)
The future of GLP-1 Peptide Mapping Using LC-MS includes automated sample preparation and Multi-Attribute Methods (MAM). MAM allows simultaneous monitoring of sequence identity, PTMs, and impurities. Automation standardizes reduction, alkylation, and digestion steps. This reduces variability and human error. High-throughput workflows improve efficiency in quality control laboratories.
Conclusion
GLP-1 Peptide Mapping Using LC-MS is an essential analytical strategy for confirming sequence identity, detecting impurities, and supporting regulatory compliance of modern GLP-1 therapeutics. By combining multi-enzyme digestion, optimized chromatographic separation, and high-resolution mass spectrometry, laboratories can obtain reliable structural information. The approach supports impurity mapping, modification localization, and comparability studies across batches. As GLP-1 analogs continue to evolve, advanced LC-MS workflows remain critical for characterization and quality control. Continued improvements in automation and MAM will further enhance reproducibility and throughput for regulated laboratories.
Frequently Asked Questions (FAQs)
Complete sequence coverage is achieved using a multi-enzyme digestion strategy that combines Trypsin, Glu-C, and Chymotrypsin. These enzymes generate overlapping peptide fragments that compensate for cleavage sites blocked by acylation. The overlapping fragments help confirm regions that are difficult to digest using a single enzyme. This approach improves mapping accuracy and ensures reliable identification of modified residues.
Difluoroacetic acid (DFA) provides strong chromatographic separation similar to TFA while causing less signal suppression in the mass spectrometer. Reduced suppression improves sensitivity and helps detect low-level impurities. DFA also maintains sharp peak shapes and stable retention times. This balance between separation quality and MS response makes DFA suitable for peptide mapping workflows.
D-amino acid isomers are separated using stationary phases that provide selective interactions, such as phenyl-hexyl or modified C18 columns. Very shallow gradient conditions help distinguish subtle structural differences between isomers. Although D- and L-forms share identical mass, their three-dimensional structures interact differently with the column. This difference enables chromatographic separation before MS confirmation.
Higher column temperatures, typically between 60°C and 80°C, improve the separation of lipidated peptides. Elevated temperature reduces mobile phase viscosity and enhances mass transfer for large hydrophobic fragments. This results in sharper peaks and improved resolution. It also helps reduce carry-over commonly observed with acylated GLP-1 analogs.
Carry-over can be reduced by using C4 columns instead of highly retentive C18 phases. Strong wash steps with high organic solvent concentrations also help remove residual peptides. Bioinert LC systems and low-bind consumables further reduce adsorption to surfaces. These measures improve recovery and maintain consistent performance across injections.
A 300 Å pore size allows large GLP-1 peptide fragments and lipidated chains to access the stationary phase efficiently. Smaller pore sizes restrict diffusion and lead to broader peaks. Improved accessibility enhances separation and peak shape. This is particularly important for large and hydrophobic GLP-1 analogs.
Site-specific acylation is confirmed by detecting characteristic mass shifts on modified fragments. Tandem MS generates fragment ion series that localize the modification to a specific residue. Diagnostic ions from spacer groups provide additional confirmation. Together, these signals verify the position and integrity of the acylation.
Validation of GLP-1 peptide mapping follows ICH Q6B guidelines for structural characterization. Analytical method validation requirements are defined in ICH Q2(R1). For generic peptide products, FDA API sameness expectations guide comparative studies. These frameworks ensure consistency, accuracy, and regulatory compliance.
HCD is preferred because it generates more complete b- and y-ion fragment series. These ions improve sequence confirmation across the peptide backbone. Higher energy fragmentation also supports analysis of stable acylated regions. As a result, HCD often provides better coverage than traditional CID.
Reference:
- Maheshwari, D., Badgujar, D., Kumar, G., & Sharma, N. (2026). Peptide mapping using multienzyme digestion strategies integrated with LC-HRMS workflow: A case study. Journal of Peptide Science, 32(3), e70089. https://doi.org/10.1002/psc.70089
- Kristensen, D. B., Kjeldsen, F., & Højrup, P. (2022). Optimized multi-attribute method workflow addressing missed cleavages in peptide mapping for improved robustness and data interpretation. Journal of the American Society for Mass Spectrometry, 33(12), 2172–2183. https://pmc.ncbi.nlm.nih.gov/articles/PMC9753059/
- U.S. Pharmacopeia (USP). (2025). GLP-1 solutions: Empowering quality & confidence. U.S. Pharmacopeia. https://www.usp.org/sites/default/files/usp/document/our-work/biologics/GLP-1%20Flyer.pdf
- Beijing ZhiTai Biopharmaceutical Technology Co., Ltd. (2020). Purification method of glucagon-like peptide-1 analogue (CN111718407A). Google Patents. https://patents.google.com/patent/CN111718407A/en



