Introduction:
GLP-1 Peptide Sequencing Challenges have become increasingly significant as modern peptide therapeutics incorporate non-natural amino acids and structural modifications to improve stability, efficacy, and half-life. While these enhancements benefit drug performance, they introduce substantial analytical complexity during sequencing and characterization.
At ResolveMass Laboratories Inc., extensive experience in GLP-1 peptide sequencing and analytical characterization of GLP-1 peptide drugs has shown that traditional workflows often fall short when dealing with modified GLP-1 analogs. This blog explores the major technical and analytical hurdles—and how to overcome them.
Share via:
Summary :
- GLP-1 Peptide Sequencing Challenges arise mainly due to non-natural amino acids, chemical modifications, and complex structures.
- Modified GLP-1 analogs often resist standard enzymatic digestion and fragmentation.
- Advanced techniques like LC-MS characterization of GLP-1 peptides and HRMS are essential.
- Data interpretation becomes complex due to isobaric residues and unexpected modifications.
- Regulatory requirements for GLP-1 peptide characterization increase analytical rigor.
- Expertise in peptide chemistry and mass spectrometry is critical for reliable results.
1: What makes GLP-1 Peptides difficult to Sequence?
GLP-1 peptides are difficult to sequence because they frequently contain non-natural amino acids, chemical modifications, and structural constraints that interfere with standard analytical workflows such as enzymatic digestion and LC-MS/MS analysis.
Key Complexity Drivers
- Non-natural amino acids (e.g., Aib, D-amino acids)
These are not well-represented in standard databases and often produce unpredictable fragmentation patterns, making identification challenging. - Post-synthetic modifications (lipidation, PEGylation)
Modifications increase molecular weight and heterogeneity, leading to ionization suppression and complex MS signals. - Disulfide bridges and secondary structures
These structural features restrict peptide unfolding and limit access for enzymatic cleavage, reducing sequence coverage. - Protease resistance
Modified GLP-1 analogs are designed to resist enzymatic degradation, which also makes controlled digestion for sequencing more difficult.
Why This Matters in Practice
These combined factors:
- Disrupt enzymatic digestion efficiency
- Complicate MS/MS fragmentation patterns
- Reduce sequence coverage and confidence
- Increase reliance on advanced techniques and expert interpretation
As a result, sequencing GLP-1 peptides requires specialized analytical strategies, including multi-enzyme digestion, high-resolution mass spectrometry, and de novo sequencing approaches to achieve accurate characterization.
Challenge 1: Non-Natural Amino Acids
Non-natural amino acids complicate GLP-1 peptide sequencing because they are not recognized by standard protein databases and often do not follow predictable MS/MS fragmentation rules.
These challenges are commonly addressed in advanced GLP-1 analog peptide sequencing workflow approaches.
Why This Is a Problem
- No reference spectra in databases
Most bioinformatics tools rely on known amino acid libraries. Non-natural residues (e.g., Aib, D-amino acids) are absent, leading to identification gaps. - Altered fragmentation behavior
These residues can disrupt typical b/y ion formation, resulting in unusual or incomplete fragmentation patterns. - Misidentification or missed identification
Standard search algorithms may incorrectly assign residues or fail to detect modified regions altogether.
Practical Impact
| Issue | Impact on Sequencing |
|---|---|
| Unknown residues | Incorrect sequence assignment |
| Fragmentation variability | Poor MS/MS interpretation |
| Database mismatch | Increased manual analysis |
Solution Approaches
- Custom database creation
- Incorporating expected non-natural amino acids into search libraries improves identification accuracy. De novo sequencing algorithms
- Enables sequence determination without relying on databases, especially useful for unknown modifications. High-resolution mass spectrometry (HRMS)
- Provides precise mass measurements to distinguish subtle differences and confirm modified residues.
Challenge 2: Chemical Modifications (Lipidation, Pegylation)
Chemical modifications such as lipidation and PEGylation make GLP-1 peptide sequencing more difficult by increasing molecular weight, introducing heterogeneity, and disrupting standard detection and fragmentation behavior in LC-MS/MS workflows.
These modifications are also critical in GLP-1 peptide stability analytical methods and long-term drug performance evaluation.
Common Modifications in GLP-1 Analogs
- Fatty acid conjugation (for half-life extension)
Enhances albumin binding but adds hydrophobic complexity and mass variability. - PEG chains (PEGylation)
Introduces polydispersity, resulting in multiple mass variants and broad signal distributions. - Acylation or amidation
Alters peptide charge, stability, and fragmentation characteristics.
Analytical Challenges
- Broad peaks in chromatography
Heterogeneous modifications (especially PEGylation) lead to poor peak resolution and difficult separation. - Suppressed ionization in MS
Lipidated or PEGylated peptides may ionize inefficiently, reducing sensitivity and signal intensity. - Complex fragmentation patterns
Modifications can dominate fragmentation pathways, masking critical sequence ions.
Best Practices
- Use LC-MS with optimized gradients
Tailored chromatographic conditions improve separation of modified variants and reduce co-elution. - Apply top-down and middle-down approaches
These strategies preserve modification information and help localize changes more effectively than bottom-up alone. - Perform orthogonal confirmation (NMR, enzymatic mapping)
Complementary techniques validate modification sites and ensure structural accuracy.
Challenge 3: Enzymatic Digestion Limitations
Modified GLP-1 peptides often resist enzymatic digestion, resulting in incomplete or non-specific cleavage that limits accurate sequencing and characterization.
Proper digestion strategies are essential for peptide sequencing of GLP-1 drugs and achieving full sequence coverage.
Why Digestion Fails
- Steric hindrance from modifications
Bulky groups such as lipid chains or PEG moieties block enzyme access to cleavage sites. - Altered enzyme recognition sites
Non-natural amino acids or chemical modifications can disrupt the specific sequences that proteases recognize. - Stable secondary structures
Folded conformations reduce accessibility, preventing efficient enzymatic interaction.
Consequences
- Incomplete peptide mapping
Key regions of the peptide remain uncharacterized due to missing fragments. - Large undigested fragments
Poor cleavage leads to oversized peptides that are difficult to analyze by LC-MS/MS. - Reduced sequence coverage
Gaps in fragmentation data lower confidence in sequence confirmation.
Mitigation Strategies
- Use multiple enzymes (Trypsin, Chymotrypsin, Glu-C)
Combining proteases with different specificities improves overall digestion efficiency. - Optimize digestion conditions
Adjust pH, temperature, incubation time, and denaturing agents to enhance enzyme accessibility. - Apply chemical cleavage methods
Non-enzymatic approaches (e.g., acid hydrolysis or CNBr cleavage) can target resistant regions.
Overcoming enzymatic digestion limitations requires a strategic, multi-enzyme and condition-optimized approach to achieve comprehensive sequence coverage of modified GLP-1 peptides.
Challenge 4: Complex MS/MS Fragmentation Patterns
Modified GLP-1 peptides produce complex and often unpredictable MS/MS fragmentation patterns, making accurate sequence interpretation significantly more challenging.
Advanced fragmentation techniques are a core part of LC-MS characterization of GLP-1 peptides workflows.
Fragmentation Issues
- Suppressed b/y ions
Chemical modifications and non-natural residues can reduce the formation of standard b and y ions, which are critical for sequence identification. - Dominance of internal fragments
Instead of clean terminal fragments, spectra may be dominated by internal fragments that are harder to interpret and assign. - Neutral losses masking key signals
Common neutral losses (e.g., water, ammonia, or modification-specific losses) can obscure important sequence-specific ions.
Advanced Solutions
- Use higher-energy collisional dissociation (HCD)
Generates richer fragmentation spectra and improves detection of sequence ions. - Combine with electron-transfer dissociation (ETD)
Preserves labile modifications and provides complementary fragmentation (c/z ions). - Employ hybrid fragmentation techniques
Combining HCD, ETD, and other methods enhances overall sequence coverage and accuracy.
Addressing complex MS/MS fragmentation requires a multi-fragmentation strategy and expert-level data interpretation to reliably decode modified GLP-1 peptide sequences.
Challenge 5: Isobaric and Similar Mass Residues
Isobaric amino acids (same mass) create ambiguity in sequence assignment.
Resolving these ambiguities is crucial for GLP-1 peptide impurity characterization and structural validation.
Examples
- Leucine vs Isoleucine
These amino acids have identical masses, making them indistinguishable in standard MS analysis. - Modified vs unmodified residues
Certain chemical modifications can produce mass shifts that overlap with naturally occurring residues.
Why It Matters
- Misinterpretation of sequence
Incorrect residue assignment can lead to inaccurate structural characterization. - Regulatory non-compliance risk
Incomplete or incorrect identification of peptide structure can impact regulatory submissions and product approval.
Resolution Techniques
- Ion mobility spectrometry (IMS)
Separates ions based on shape and size, helping distinguish structurally similar residues. - Advanced fragmentation strategies
Techniques like HCD and ETD can generate unique fragment ions that help differentiate isobaric residues. - Orthogonal analytical confirmation
Complementary methods such as NMR or enzymatic mapping provide additional structural validation.
Challenge 6: Data Interpretation and Bioinformatics
Data interpretation in GLP-1 peptide sequencing is highly complex due to unknown modifications, large high-resolution datasets, and limited capability of standard bioinformatics tools.
Advanced tools and expertise are key in analytical characterization of GLP-1 peptide drugs.
Key Issues
- Large datasets from high-resolution MS
Advanced instruments generate massive volumes of spectral data, making processing and interpretation time-consuming. - Unknown or unexpected modifications
Non-natural amino acids and novel chemical modifications are often not included in standard databases, complicating identification. - Software limitations
Many peptide sequencing tools are optimized for natural peptides and struggle with modified or synthetic analogs.
Required Expertise
- Manual spectral interpretation
Experts must validate and interpret complex MS/MS spectra beyond automated outputs. - Custom algorithm development
Tailored bioinformatics workflows are often needed to incorporate non-standard residues and modifications. - Deep understanding of peptide chemistry
Knowledge of fragmentation behavior, modification chemistry, and analytical techniques is essential for accurate conclusions.
Successfully overcoming bioinformatics challenges requires combining advanced computational tools with expert-driven analysis to ensure accurate and reliable GLP-1 peptide sequencing results.
Challenge 7: Regulatory Expectations
Regulatory agencies require comprehensive structural characterization of GLP-1 peptides, significantly increasing the analytical rigor and complexity of sequencing workflows.
Meeting these expectations aligns with regulatory requirements for GLP-1 peptide characterization.
Requirements Include
- Full sequence confirmation
Complete and unambiguous identification of the peptide’s amino acid sequence, including non-natural residues. - Identification of all modifications
Precise localization and characterization of chemical modifications such as lipidation, PEGylation, or acylation. - Impurity and degradation profiling
Detection, identification, and quantification of related impurities and degradation products.
Impact on Sequencing
- Need for multiple orthogonal techniques
No single method is sufficient—regulatory compliance requires combining LC-MS/MS, HRMS, NMR, and other analytical tools. - Extensive documentation
Detailed reporting, method validation, and traceability are mandatory for submissions. - High reproducibility standards
Analytical methods must demonstrate consistency, robustness, and reliability across multiple runs and conditions.
Meeting regulatory expectations demands a highly integrated analytical strategy, combining advanced instrumentation, validated methods, and deep scientific expertise to ensure accurate and compliant GLP-1 peptide sequencing.
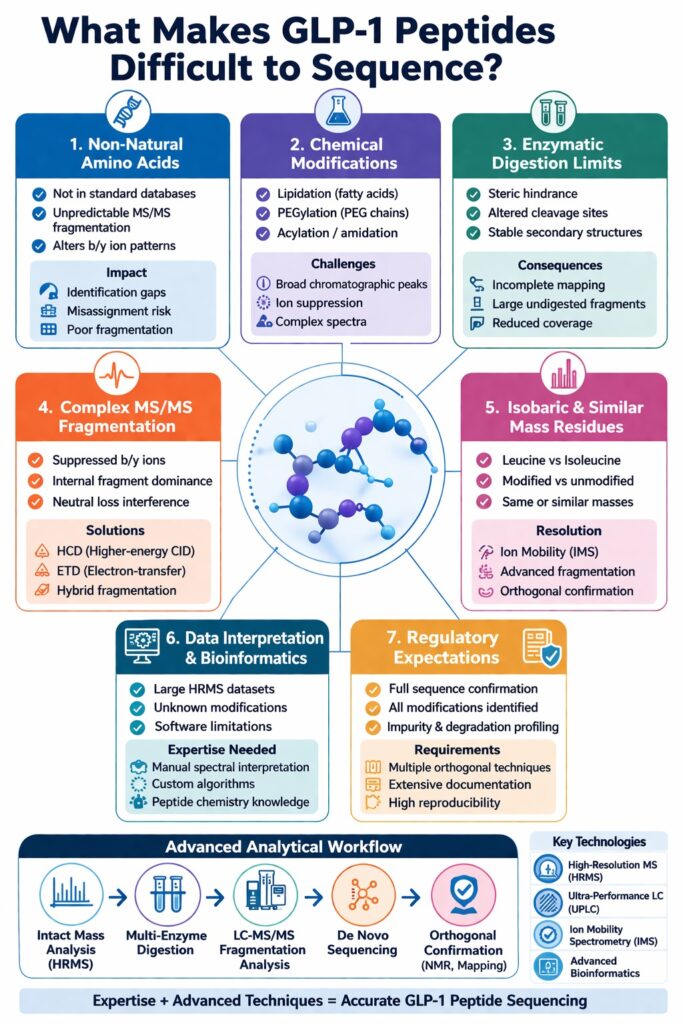
2: Advanced Analytical Strategies to Overcome Challenges
Accurate GLP-1 peptide sequencing requires combining multiple advanced analytical techniques, as no single method can fully resolve the complexity introduced by non-natural amino acids and chemical modifications.
A structured approach like the GLP-1 analog peptide sequencing workflow ensures high-confidence results.
Recommended Workflow
A robust, stepwise analytical workflow ensures comprehensive characterization:
- Intact Mass Analysis (HRMS)
Confirms the overall molecular weight and detects major modifications or mass shifts at the intact peptide level. - Enzymatic Digestion (Multi-Enzyme Approach)
Uses complementary proteases (e.g., Trypsin, Chymotrypsin, Glu-C) to improve sequence coverage and overcome digestion resistance. - LC-MS/MS Fragmentation Analysis
Provides detailed sequence information through fragmentation patterns, enabling identification of amino acid order and modification sites. - De Novo Sequencing
Determines peptide sequence without relying on databases—critical for identifying non-natural amino acids and unknown modifications. - Orthogonal Confirmation (NMR, Edman Degradation if applicable)
Validates sequence and structural features using complementary techniques for higher confidence and regulatory compliance.
Technology Stack
- High-resolution mass spectrometry (HRMS)
Enables accurate mass measurement and detection of subtle differences in modified peptides. - Ultra-performance liquid chromatography (UPLC)
Provides high-efficiency separation of complex peptide mixtures and modified variants. - Advanced bioinformatics tools
Supports data processing, de novo sequencing, and integration of custom modifications into analysis workflows.
A multi-technique, integrated strategy is essential to successfully overcome GLP-1 peptide sequencing challenges and deliver reliable, high-confidence analytical results.
3: Why Expertise matters in GLP-1 Peptide Sequencing
Sequencing modified GLP-1 peptides demands specialized expertise, advanced instrumentation, and tailored method development to accurately resolve complex structures and modifications.
Modern GLP-1 analogs are far more sophisticated than standard peptides. The presence of non-natural amino acids, lipidation, PEGylation, and structural constraints means that generic analytical approaches often fail. This is where deep scientific expertise becomes critical—not just in operating instruments, but in designing the right strategy, interpreting complex data, and ensuring regulatory compliance.
Partnering with a specialized CRO for GLP-1 peptide characterization ensures accuracy, compliance, and efficiency.
Explore dedicated GLP-1 peptide sequencing CRO services for end-to-end analytical support.
Key Areas where Expertise makes a Difference
- Handling complex peptide modifications
Expert scientists understand how different modifications affect ionization, fragmentation, and digestion, enabling accurate identification and localization. - Developing customized analytical workflows
No single method fits all GLP-1 analogs. Tailored workflows combining LC-MS/MS, HRMS, and orthogonal techniques are essential for reliable sequencing. - Advanced data interpretation
Interpreting complex MS/MS spectra, especially with unknown modifications, requires manual validation and deep knowledge of peptide chemistry. - Ensuring regulatory compliance
Experts design studies that meet global regulatory expectations, including full sequence confirmation, impurity profiling, and reproducibility.
The ResolveMass Advantage
At ResolveMass Laboratories Inc., the approach is built on:
- Proven experience in complex peptide characterization
- Strong capability in method development and optimization
- Focus on high-quality, regulatory-compliant data delivery
Why it matters for Pharmaceutical Development
- Reduces risk of sequence misidentification
- Accelerates drug development timelines
- Ensures data integrity for regulatory submissions
- Improves confidence in product quality and consistency
Ultimately, expertise is the deciding factor in successfully overcoming GLP-1 peptide sequencing challenges and delivering accurate, reproducible, and compliant analytical results.
Conclusion:
GLP-1 Peptide Sequencing Challenges are driven by the increasing complexity of modern peptide therapeutics, especially those containing non-natural amino acids and chemical modifications. These challenges impact every stage—from digestion and detection to data interpretation and regulatory compliance.
From peptide sequencing of GLP-1 peptide to impurity profiling and regulatory compliance, each step requires advanced analytical strategies and deep expertise.
Successfully overcoming these hurdles requires a combination of:
- Specialized CRO expertise
- Advanced analytical techniques
- Integrated workflows
Frequently Asked Questions:
GLP-1 Peptide Sequencing Challenges refer to the analytical difficulties in determining the exact amino acid sequence and modifications of GLP-1 analogs. These challenges arise due to non-natural amino acids, chemical modifications, and complex structures that interfere with standard LC-MS/MS workflows.
Non-natural amino acids are difficult to analyze because they are not present in standard protein databases and often exhibit unpredictable fragmentation patterns. This makes automated identification challenging and requires de novo sequencing and expert interpretation.
Chemical modifications increase molecular weight and heterogeneity, leading to ionization suppression and complex MS signals. They also alter fragmentation behavior, making it harder to identify sequence and modification sites accurately.
Modified GLP-1 peptides are designed to resist enzymatic degradation, which also makes controlled digestion difficult. Structural hindrance and altered cleavage sites often result in incomplete digestion and reduced sequence coverage.
Advanced techniques include high-resolution mass spectrometry (HRMS), LC-MS/MS, multi-enzyme digestion, de novo sequencing, and orthogonal methods like NMR. Combining these approaches ensures accurate and comprehensive peptide characterization.
Reference
- Otvos L. The latest trends in peptide drug discovery and future challenges. Expert Opinion on Drug Discovery. 2024 Aug 2;19(8):869-72.https://www.tandfonline.com/doi/abs/10.1080/17460441.2024.2365969
- Panda SS. Peptides and Amino Acids in Drug Development: Where We Stand and Where We Must Go. Biomedicines. 2026 Mar 16;14(3):678.https://www.mdpi.com/2227-9059/14/3/678
- Pereira AJ, de Campos LJ, Xing H, Conda-Sheridan M. Peptide-based therapeutics: challenges and solutions. Medicinal Chemistry Research. 2024 Aug;33(8):1275-80.https://link.springer.com/article/10.1007/s00044-024-03269-1
- Ding Y, Ting JP, Liu J, Al-Azzam S, Pandya P, Afshar S. Impact of non-proteinogenic amino acids in the discovery and development of peptide therapeutics. Amino acids. 2020 Sep;52(9):1207-26.https://link.springer.com/article/10.1007/s00726-020-02890-9
- Lombardi L, Genio VD, Albericio F, Williams DR. Advances in peptidomimetics for next-generation therapeutics: strategies, modifications, and applications. Chemical reviews. 2025 Jul 23;125(15):7099-166.https://pubs.acs.org/doi/abs/10.1021/acs.chemrev.4c00989
- Ma B, Fuhrmann J, Henriksen H, Khojasteh SC, Li W, Liu J, Plise E, Yu Q, Cheruzel L. Overcoming challenges in the metabolism of peptide therapeutics: strategies and case studies for clinical success. Journal of Medicinal Chemistry. 2025 Dec 5;68(24):25689-707.https://pubs.acs.org/doi/abs/10.1021/acs.jmedchem.5c02276
- Oeller M, Kang RJ, Bolt HL, Gomes dos Santos AL, Weinmann AL, Nikitidis A, Zlatoidsky P, Su W, Czechtizky W, De Maria L, Sormanni P. Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids. Nature Communications. 2023 Nov 17;14(1):7475.https://www.nature.com/articles/s41467-023-42940-w

