Introduction:
The Lanreotide Sameness Study is a comprehensive analytical exercise designed to prove that a generic lanreotide product is identical to the Reference Listed Drug (RLD) in sequence, structure, and purity. Unlike simple small molecules, peptides require detailed characterization. Regulatory guidance demands that the generic lanreotide matches the innovator exactly in amino acid sequence, post-translational modifications (e.g. disulfide bonds), and impurity profile. Lanreotide is a synthetic cyclic octapeptide analog of somatostatin, and its complex structure (including a critical disulfide bridge) makes rigorous analysis essential. ResolveMass Laboratories Inc. brings specialized expertise and validated methods to this challenge – our Montreal-based team has performed 40+ peptide ANDA projects using advanced LC–MS and NMR, ensuring audit-ready, FDA-compliant data.
Summary:
- Comprehensive multi-technique approach: A Lanreotide Sameness Study uses LC–MS/MS (HRMS) peptide sequencing, enzymatic peptide mapping, and orthogonal methods (intact mass, NMR) to confirm the generic matches the reference product in structure and purity.
- Peptide sequencing: High-resolution MS/MS data are used to verify the complete amino acid sequence, including cyclic disulfide bonds, of lanreotide. Fragment ions must exactly match the theoretical sequence (mass error <2 ppm) for identity confirmation.
- Peptide mapping fingerprint: Enzymatic digestion (e.g. trypsin, chymotrypsin) followed by LC–MS comparison generates a chromatographic peptide fingerprint. The test and RLD must exhibit identical peptide peaks (same retention times and masses).
- Impurity profiling by HRMS: High-resolution LC–MS analysis detects and quantifies process or degradation impurities (truncated peptides, oxidation variants, deamidation, D/L isomers). Observed impurities must mirror the reference profile, with no new impurities above qualification thresholds. Typical detection limits are ~0.1% for truncations and ~0.2% for oxidized variants.
- Intact mass analysis: The intact lanreotide peptide is measured by HRMS to confirm its total molecular weight matches the RLD (monoisotopic mass ~1096.57 Da) within tight error (typically ≤5 ppm).
- NMR confirmation: 1D and 2D NMR provide orthogonal structural confirmation (e.g. verifying disulfide connectivity, ring conformation and absence of unexpected modifications). Key NMR signals (e.g. aromatic or cysteine peaks) must match between test and reference.
- Regulatory compliance: All methods and data handling are performed in a GLP/GMP-ready framework (21 CFR Part 11 compliant systems) and aligned to FDA/ICH peptide guidance. A final comparative report demonstrates that the generic lanreotide is analytically indistinguishable from the Reference Listed Drug (RLD).
1: Analytical Strategy Overview (Multi-Technique Approach)
ResolveMass employs a multi-technique analytical platform to cover all critical quality attributes of lanreotide. Key components include: high-resolution LC–MS/MS for exact sequence and mass, enzymatic peptide mapping for fingerprinting, LC–MS impurity profiling for degradation and process variants, intact mass measurement for overall molecular weight, and NMR for conformational/structural confirmation.
Figure: Schematic LC–MS/MS workflow used in peptide sequencing and mapping. Peptides are separated by liquid chromatography, ionized, and fragmented in MS/MS for sequence information.
Each technique is chosen to address a specific attribute. For example, tandem MS fragments peptides to confirm amino acid order, while HRMS ensures precise mass measurement. The enzymes used for digestion are selected to generate overlapping peptides (e.g. trypsin, chymotrypsin), ensuring 100% sequence coverage. Chromatographic methods (typically reversed-phase HPLC with water/acetonitrile gradients) separate fragments and impurities. All analytical data are collected under 21 CFR Part 11‐compliant conditions for traceability and audit readiness.
2: Peptide Sequencing by HRMS/MS
2.1 Objective
Objective: Obtain unequivocal amino acid sequence of the generic lanreotide and confirm its cyclic disulfide bond, matching the RLD. HRMS/MS must verify that the generic has the exact same peptide sequence, length, and post-translational modifications as the reference. Even a single amino acid difference would violate sameness criteria.
2.2 Structural Considerations
Lanreotide’s structure is a cyclic octapeptide featuring an internal disulfide bond. This requires fragmentation techniques that preserve or reveal the disulfide connectivity. We account for lanreotide’s unique modifications (e.g. d-amino acids, methylations) in our analysis. The disulfide bond is probed by comparing reduced versus non-reduced peptide masses, and by characteristic fragment patterns.
2.3 Enzymatic Digestion Strategy
To sequence the peptide, the cyclic structure is first linearized by reducing the disulfide bond (if needed) or by cleaving with specific proteases. We typically use trypsin and chymotrypsin to generate overlapping peptides. The choice of enzymes ensures each peptide fragment is manageable by MS/MS and provides full coverage of the sequence. Any cleavage pattern is carefully planned to avoid missing the N- or C-terminus.
2.4 Experimental Procedure
Digest samples are analyzed by nano-LC coupled to a high-resolution mass spectrometer (e.g. Orbitrap) in positive-ion mode. Peptides are ionized by electrospray and subjected to collision-induced dissociation (CID or HCD) in the MS/MS cell. The resulting fragment ions (b and y ions) are recorded with high mass accuracy (often <5 ppm). All runs include the RLD material as a direct control. Data are collected and processed using validated software, ensuring reliable identification of each fragment.
2.5 Data Analysis
MS/MS spectra are compared to the theoretical spectra of lanreotide. We require that all expected fragments of the peptide sequence are observed, with mass errors below ~2 ppm. Software automatically matches fragment ions, and manual review ensures any ambiguities (e.g. isobaric residues) are resolved. Disulfide bond confirmation is done by observing the mass of the reduced peptide and by verifying the continuity of sequential fragments across the original bridge.
2.6 Key Observations / Acceptance Criteria
The generic passes the sequencing test when it meets all of the following: 100% sequence identity with the RLD (no missing or substituted residues), correct cyclic disulfide bond pattern, and fragment ion masses within acceptance limits (<2 ppm). Chromatographic peak retention times of the peptide and major fragments should also coincide with the reference. All data are archived in a 21 CFR Part 11–compliant system to ensure data integrity.
3: Peptide Mapping (Comparative Fingerprinting)
3.1 Objective
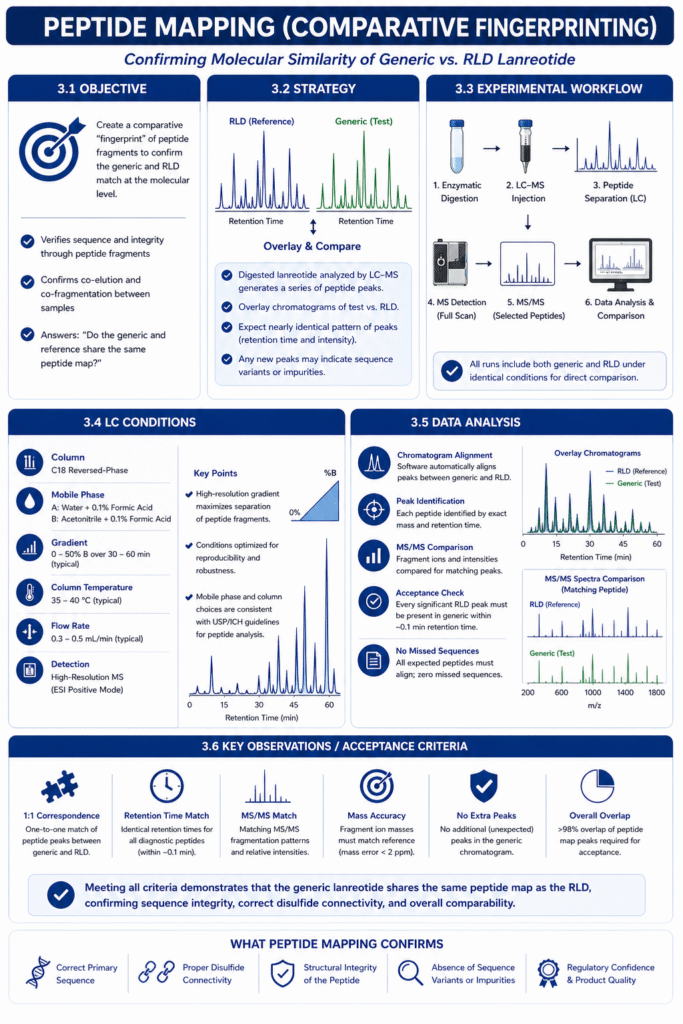
Objective: Create a comparative “fingerprint” of peptide fragments to confirm the generic and RLD match at the molecular level. Peptide mapping verifies sequence and integrity by showing that digested peptide fragments co-elute and co-fragment between samples.
3.2 Strategy
The digested lanreotide sample is analyzed by LC–MS, producing a series of peptide peaks. We overlay the chromatograms of the test versus RLD and expect a nearly identical pattern of peaks (in retention time and intensity). Any new peaks could indicate sequence variants or impurities. This approach directly answers, “Do the generic and reference share the same peptide map?”.
3.3 Experimental Workflow
After digestion, peptides are injected onto an LC–MS system. A typical setup uses a C18 reversed-phase column and a water/acetonitrile gradient (both with 0.1% formic acid). The eluting peptides are detected by MS, and key fragments are further analyzed by MS/MS if needed. All runs include both generic and RLD under identical conditions for direct comparison.
3.4 LC Conditions
We use a high-resolution LC gradient to maximize separation of peptide fragments. For example, a 0–50% acetonitrile gradient over 30–60 minutes often resolves the lanreotide peptides. Column temperature and flow rate are optimized for reproducibility. Mobile phase and column choices are documented and consistent with USP/ICH guidelines for peptide analysis.
3.5 Data Analysis
Chromatograms of generic and RLD digests are compared peak-by-peak. Each observed peptide is identified by its exact mass and retention time. Software automatically aligns peaks, and inspectors verify that every significant peak in the RLD map appears in the generic at the same retention time (within ~0.1 min). MS/MS spectra of matching peaks are also compared: fragment ions and intensities should be congruent. We look for zero “missed” sequences – all expected peptides must align.
3.6 Key Observations / Acceptance Criteria
Acceptance requires a 1:1 correspondence between generic and RLD peptides. Specifically: identical retention times for all diagnostic peptides, matching MS/MS fragmentation patterns, and no extra peaks in the generic trace. Fragment ion masses must match reference spectra (mass error <2 ppm). Retention time reproducibility also checks disulfide connectivity in cyclic regions. In practice, we demand >98% overlap of peptide map peaks.
4: Impurity Profiling by HRMS
4.1 Objective
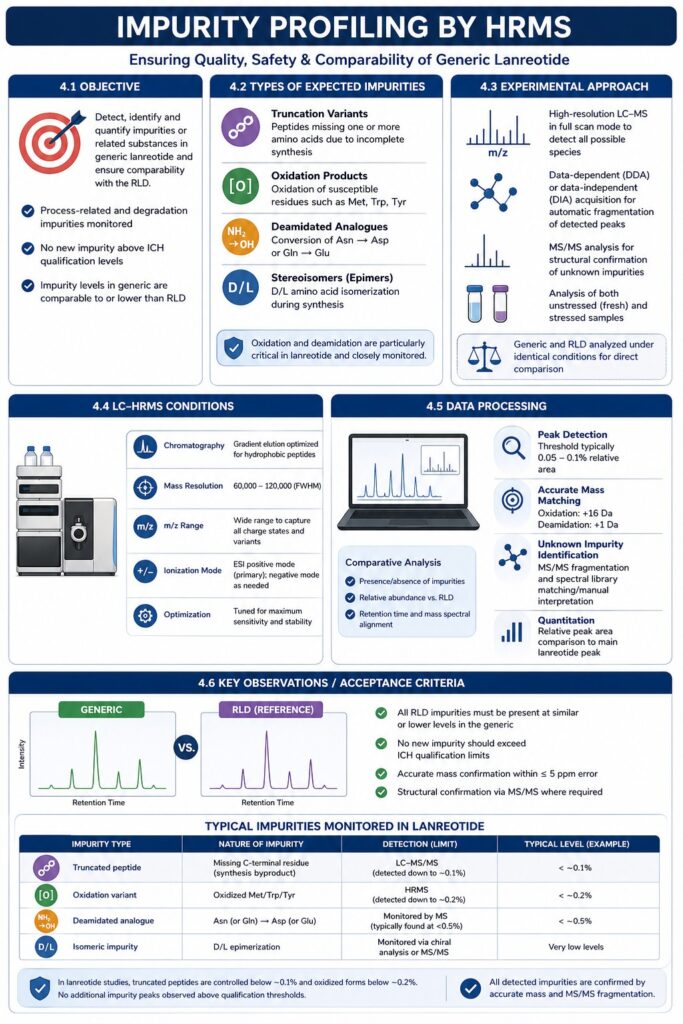
Objective: Detect and quantify any impurities or related substances in the generic lanreotide and ensure they are comparable to (or fewer than) those in the RLD. This includes process-related byproducts and degradation products. The goal is to confirm no new impurity above ICH qualification levels appears in the generic.
4.2 Types of Expected Impurities
In synthetic peptides like lanreotide, common impurities include: truncation variants (peptides missing one or more amino acids), oxidized or reduced forms (e.g. Met or Trp oxidation), deamidated analogues (Asn→Asp), and stereoisomers (D/L epimers). Each of these can arise during synthesis, formulation, or storage. For lanreotide specifically, oxidation and deamidation of susceptible residues are closely monitored.
4.3 Experimental Approach
We use high-resolution LC–MS in full scan mode, often with a fast MS/MS screening of unknown peaks. Data-dependent acquisition (or DIA) can be employed so that any peak above the noise is automatically fragmented for identification. Both fresh and stressed samples are analyzed. The test and RLD are run under identical gradient conditions to directly compare impurity profiles.
4.4 LC–HRMS Conditions
A gradient optimized for hydrophobic peptides is used (similar to the mapping workflow). The mass spectrometer is set to high resolution (e.g. 60–120k), covering a wide m/z range to catch all species. Electrospray ionization parameters are tuned to maximize sensitivity for lanreotide and its variants. Where applicable, we may run analyses in both positive and negative mode to catch different chemistries.
4.5 Data Processing
Impurity peaks are extracted and compared between samples. We use software to find peaks present in the generic above a certain threshold (e.g. 0.05–0.1%). Each candidate impurity is identified by its accurate mass: known modifications (oxidation: +16 Da, deamidation: +1 Da) are matched, and unknowns are investigated by MS/MS. Quantitation is performed by comparing peak areas to the main lanreotide peak.
4.6 Key Observations / Acceptance Criteria
The generic’s impurity profile should mirror the reference’s. That means: all impurities seen in the RLD must be present (at similar or lower levels) in the test, and no new impurity should exceed qualification limits. Using the example list from ResolveMass studies:
| Impurity Type | Nature of Impurity | Detection (Limit) |
|---|---|---|
| Truncated peptide | Missing C-terminal residue (synthesis byproduct) | LC–MS/MS (detected down to ~0.1%) |
| Oxidation variant | Oxidized Met/Trp/Tyr | HRMS (detected down to ~0.2%) |
| Deamidated analogue | Asn (or Gln) → Asp (or Glu) | Monitored by MS; typically found at <0.5% |
| Isomeric impurity | D/L epimerization | Monitored via chiral analysis or MS/MS fragment patterns |
For example, in our lanreotide studies, truncated peptides are controlled below ~0.1% and oxidized forms below ~0.2%. Absence of extra peaks above these levels is required. All impurity masses are confirmed by exact mass (≤5 ppm error) and, when needed, by MS/MS fragmentation.

5: Intact Mass Analysis
5.1 Objective
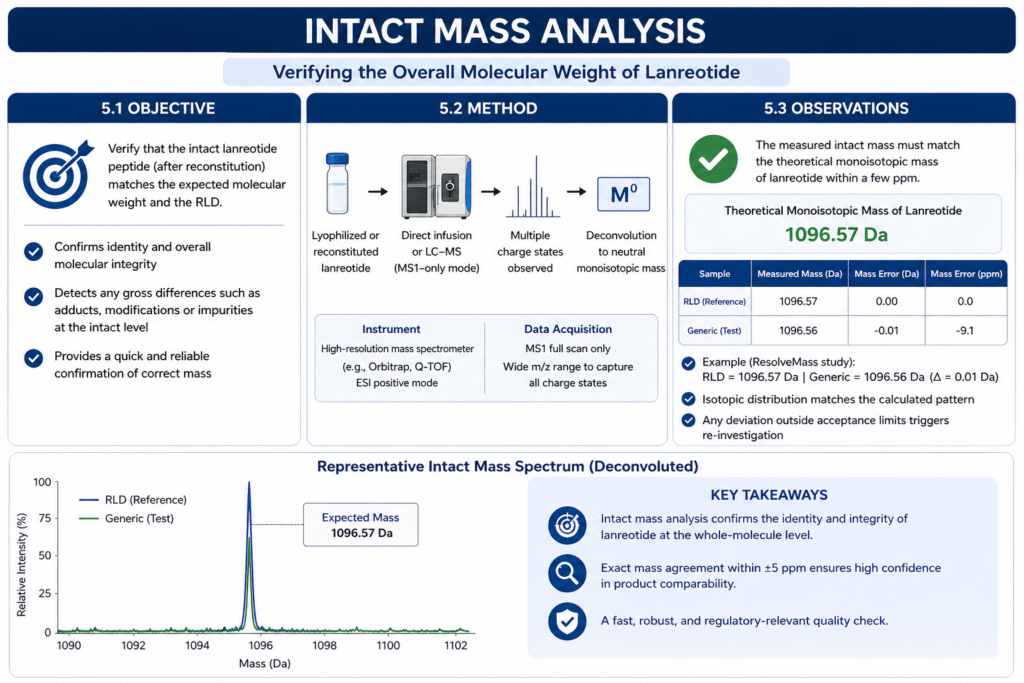
Objective: Verify the overall molecular weight of lanreotide to confirm that the intact peptide (after reconstitution) matches the expected mass and the RLD. This is a quick check for any gross differences (e.g. adducts, modifications).
5.2 Method
The lyophilized peptide (or reconstituted formulation) is directly infused or run by LC–MS in MS1-only mode on a high-resolution instrument. We detect multiple charge states of the full peptide and deconvolute to the neutral monoisotopic mass.
5.3 Observations
The measured intact mass must be equal to the theoretical monoisotopic mass of lanreotide (1096.57 Da) within a few ppm. For example, ResolveMass found the reference lanreotide at 1096.57 Da and the generic at 1096.56 Da (a 0.01 Da difference). We also check isotopic distribution and confirm it matches the calculated pattern. Any deviation would trigger re-investigation.
6: NMR Characterization (Orthogonal Confirmation)
6.1 Objective
Objective: Provide an orthogonal structural check of lanreotide’s higher-order features and purity. NMR helps confirm disulfide bonds, ring conformation, and identify any unexpected chemical modifications. It also offers a quantitative purity measure via peak integration.
6.2 Techniques
We acquire 1D ^1H NMR (and ^13C NMR if needed) plus 2D spectra (e.g. COSY, HSQC, TOCSY). The 1D ^1H spectrum of lanreotide shows characteristic aromatic and aliphatic resonances for the eight residues. 2D spectra help assign connectivities and verify sequence continuity. Quantitative NMR (qNMR) can also be used to assess overall purity by integrating known singlet peaks against an internal standard.
6.3 Key Focus Areas
We focus on identifying signals unique to lanreotide’s cyclic structure and verifying the disulfide bond. For example, cysteine Hβ protons and aromatic rings (Phe, Trp) have distinctive chemical shifts. Matching these peaks between test and reference confirms identical structure. We also inspect for extra peaks that would indicate impurities (non-peptidic additives or breakdown products). NMR can even detect lanreotide’s nanotube self-association state under certain conditions.
Figure: An NMR spectrometer, such as this 11.7 Tesla instrument, is used for peptide structural analysis. High-field NMR provides detailed information on peptide bonds and conformation.
This NMR instrument was used to acquire lanreotide spectra. Key acceptance criteria include: identical chemical shift patterns (no new peaks) between generic and RLD; and quantitative peak integration within experimental error. For instance, 2D NMR should show the expected number of cross-peaks for an octapeptide with one disulfide. These orthogonal data complement the MS findings.
6.4 Acceptance Criteria
The generic lanreotide is accepted by NMR if all major peaks match the reference (within ~0.02 ppm), and no extra resonances are observed. Integration of peptide signals should indicate >98% purity. In practice, we require >95% overlap in 2D correlation maps. Combined with the LC–MS data, matching NMR provides high confidence in structural equivalence.
7: Comparative Sameness Assessment (RLD vs Test)
After all analyses, data from generic and RLD are compared side-by-side to demonstrate sameness. As illustrated in ResolveMass case studies, key parameters such as monoisotopic mass, purity, and retention times must align. For example, one study showed:
| Parameter | Reference Lanreotide | Generic Candidate | Analytical Tool |
|---|---|---|---|
| Monoisotopic Mass | 1096.57 Da | 1096.56 Da | HRMS |
| RP-HPLC Purity | 99.1% | 99.0% | HPLC |
| LC Retention Time | 9.82 min | 9.80 min | LC–MS |
| CD Spectral Overlap | – | ≥98% overlap | Circular Dichroism |
These matched results “confirm analytical sameness as required by the FDA”. In practice, our final report compiles tables and plots (MS spectra, chromatograms, NMR overlays) that clearly demonstrate equivalence. Any minor variations must be justified (e.g. batch variability). Overall, the generic must be indistinguishable from the RLD by all measured criteria.
8: Regulatory Considerations for ANDA
Regulators expect a thorough characterization package for lanreotide generics. The FDA’s product-specific guidance explicitly notes that a Q1/Q2 equivalent lanreotide (same composition and dosage form) may qualify for a biowaiver if the test demonstrates “equivalent molecular, structural, and thermodynamic properties” to the RLD. In other words, analytical data must show 1:1 sameness. Current FDA guidelines (2021) emphasize orthogonal methods and detailed impurity assessment for peptides. Thus, our approach aligns with FDA/ICH expectations by covering sequence, mass, purity, structure, and stability. Providing validated, reproducible data (with audit trails) significantly de-risks the ANDA review.
Conclusion:
A robust Lanreotide Sameness Study is essential for a successful ANDA filing. By combining high-resolution mass spectrometry, peptide mapping, impurity profiling, intact mass confirmation, and NMR, we comprehensively prove that the generic lanreotide matches the innovator in every key attribute. ResolveMass Laboratories’ depth of experience, PhD-level team, and 40+ peptide ANDA project track record ensure expert execution and FDA-readiness. Our validated, compliant data package minimizes regulatory risk and streamlines approval.
Frequently Asked Questions:
Peptide mapping is an analytical technique used to compare the fragmentation pattern of peptides between a generic and reference product. In Lanreotide studies, it confirms that both products share the same amino acid sequence and structural integrity. By enzymatically digesting the peptide and analyzing fragments via LC–MS, scientists generate a unique “fingerprint.” Matching fingerprints indicate molecular equivalence. This is a critical requirement for ANDA approval.
Comparative peptide mapping ensures that the generic drug is structurally identical to the RLD at the molecular level. It helps detect even minor sequence variations, degradation, or synthesis-related differences. Regulatory agencies require this level of detail to confirm drug safety and efficacy. A high degree of overlap (>98%) strengthens the case for sameness. Without it, approval risks increase significantly.
LC–MS separates peptide fragments based on retention time and identifies them using accurate mass measurements. The LC component resolves complex mixtures, while MS detects and characterizes each fragment. MS/MS further confirms peptide identity through fragmentation patterns. Together, they provide a powerful platform for detailed comparison. This ensures both qualitative and quantitative alignment between generic and RLD.
Acceptance requires a 1:1 correspondence of peptide peaks between generic and reference products. Retention times must match within approximately 0.1 minutes. MS/MS spectra should show identical fragment ions and relative intensities. Mass accuracy must typically fall within ±2 ppm. Additionally, no unexpected peaks should appear in the generic profile.
Co-elution refers to peptide fragments from both samples appearing at the same retention time during LC separation. Co-fragmentation means these fragments produce identical MS/MS spectra, confirming structural similarity. Together, these parameters validate that the peptides are chemically and structurally identical. This is a strong indicator of sameness. Any deviation may suggest impurities or sequence differences.
Yes, peptide mapping is highly sensitive and can detect low-level impurities or sequence variants. Any new or unmatched peak in the chromatogram may indicate a modification, truncation, or impurity. MS/MS analysis helps identify the exact nature of these differences. This makes peptide mapping not just a confirmation tool, but also a diagnostic one. It is especially valuable in complex peptide drugs.
Typically, regulators expect greater than 98% overlap between peptide maps. This includes matching retention times, peak intensities, and MS/MS spectra. All critical peptides must be present with no missing sequences. Minor variations, if any, must be scientifically justified. High similarity confirms structural sameness and supports regulatory approval.
Reference
- Wang L, Chen S, Wu M, Zhou L. Comparative analysis of adverse event profiles of lanreotide and octreotide in somatostatin-responsive endocrine and neoplastic diseases. Scientific Reports. 2025 May 28;15(1):18641.https://www.nature.com/articles/s41598-025-03850-7
- Chanson P, Boerlin V, Ajzenberg C, Bachelot Y, Benito P, Bringer J, Caron P, Charbonnel B, Cortet C, Delemer B, Escobar‐Jiménez F. Comparison of octreotide acetate LAR and lanreotide SR in patients with acromegaly. Clinical Endocrinology. 2000 Nov 17;53(5):577-86.https://onlinelibrary.wiley.com/doi/abs/10.1046/j.1365-2265.2000.01134.x
- Huynh L, Cai B, Cheng M, Lax A, Lejeune D, Duh MS, Kim MK. Analysis of real-world treatment patterns, healthcare resource utilization, and costs between octreotide and lanreotide among patients with neuroendocrine tumors. Pancreas. 2019 Oct 1;48(9):1126-35.https://journals.lww.com/pancreasjournal/fulltext/2019/10000/Analysis_of_Real_World_Treatment_Patterns,.3.aspx
- Chowdhury A, Tripathi NM, Jadav R, Gour V, Purohit P, Bandyopadhyay A. On-resin synthesis of Lanreotide epimers and studies of their structure–activity relationships. RSC Medicinal Chemistry. 2024;15(8):2766-72.https://pubs.rsc.org/en/content/articlehtml/2024/md/d4md00338a
- Grimaldi M, Santoro A, Buonocore M, Crivaro C, Funicello N, Sublimi Saponetti M, Ripoli C, Rodriquez M, De Pasquale S, Bobba F, Ferrazzano L. A new approach to supramolecular structure determination in pharmaceutical preparation of self-assembling peptides: A case study of lanreotide Autogel. Pharmaceutics. 2022 Mar 20;14(3):681.https://www.mdpi.com/1999-4923/14/3/681


