Introduction:
The Liraglutide Sameness Study is essential to prove that a generic liraglutide product is molecularly identical to the reference listed drug (RLD). Under the FDA’s 505(j) pathway, an ANDA applicant must demonstrate that the active ingredient’s primary structure, chemical modifications, and impurity profile match the innovator product. Liraglutide is a 31-amino-acid GLP-1 analog with 97% sequence homology to native GLP-1, distinguished by a site-specific C16 palmitoyl fatty acid attached at Lys26 via a γ-glutamic acid linker. This complex peptide (marketed as Victoza/Saxenda) resists rapid degradation and binds albumin for prolonged action. Its modified structure and multiple potential degradation sites (e.g. Met oxidation, Asn deamidation) make sameness testing far more demanding than for small molecules.
Regulatory guidance explicitly states that for synthetic peptide ANDAs, demonstrating sameness relies heavily on analytical comparisons of impurities and sequence. In fact, the FDA treats peptides ≤40 amino acids as chemically synthesized drugs, not biologics. Thus, an ANDA must include rigorous molecular characterization. Key attributes to confirm include identical amino acid sequence and modifications, accurate intact mass, and a comparable impurity profile. As a result, a Liraglutide Sameness Study deploys multiple orthogonal techniques (mass spectrometry and NMR) to meet FDA standards. This article outlines a comprehensive analytical strategy – from sequencing and mapping to impurity profiling and NMR – that our team at ResolveMass Laboratories Inc. has successfully used to support ANDA filings with regulatory-grade data.
Summary:
- Regulatory Requirement: An ANDA for liraglutide must prove the generic’s API is identical in sequence, modification, and impurities to the reference drug.
- Multidimensional Analytics: A successful Liraglutide Sameness Study combines orthogonal methods – intact mass (LC-HRMS), peptide sequencing/mapping (LC-MS/MS), impurity profiling (HRMS), and NMR – to cover all structural levels.
- Peptide Sequencing: 100% primary sequence identity (including the Lys26 palmitoyl modification) is confirmed by MS/MS fragmentation; all expected peptide fragments must match the reference.
- Peptide Mapping: Comparative LC-MS mapping of enzymatic digests ensures complete sequence coverage (≥100%) and identifies post-translational or degradation modifications.
- Impurity Profiling: High-resolution LC-MS detects trace peptide-related impurities (degradation products, sequence variants, process impurities). The generic’s impurity profile must be qualitatively and quantitatively comparable to the RLD.
- NMR (HOS Analysis): 1D/2D NMR provides an orthogonal fingerprint of the peptide’s higher-order structure. Matching chemical shifts and cross-peaks between test and RLD confirm conformational equivalence.
- Comparative Assessment: Summary metrics (sequence match, mass accuracy, impurity levels) are tabulated to demonstrate equivalence. Regulators expect all data in a validated, CTD-compliant package.
- Expert Support: ResolveMass Laboratories’ multidisciplinary team leverages specialized peptide analysis experience to ensure your ANDA submission meets FDA expectations and avoids deficiencies.
1: Analytical Strategy Overview (Multi-Technique Approach)
A robust Liraglutide Sameness Study requires a multi-technique analytical workflow that covers every structural dimension of the peptide. In practice, this means combining:
- Intact Mass Analysis (LC-HRMS): Confirms the exact molecular weight of the peptide, verifying the full sequence plus the Lys26 palmitoylation (theoretical M ≈ 3751.20 Da).
- Peptide Sequencing (LC-MS/MS): Confirms the 100% amino acid sequence by fragmenting the digested peptide and matching b/y ions to the reference sequence.
- Peptide Mapping (Comparative Fingerprint): Generates a complete peptide map (HPLC fingerprint) for both generic and RLD under identical conditions, ensuring no sequence gap or unaccounted modification.
- Impurity Profiling (LC-HRMS): Identifies and quantifies all process- and degradation-related impurities at trace levels, comparing their profiles between generic and RLD.
- NMR Spectroscopy (1D & 2D): Provides orthogonal evidence of higher-order structure. NMR fingerprints (chemical shifts, couplings) must align between products to confirm conformational sameness.
- Forced Degradation Studies: Generating stress samples (oxidation, heat, light, hydrolysis) to verify that degradation pathways and impurities behave similarly in both samples.
These steps are guided by best practices in peptide characterization and ANDA guidance. For example, the FDA expects a combination of high-resolution mass spectrometry and complementary methods (HPLC-UV, ion chromatography, etc.) to fully characterize peptides. This orthogonal strategy ensures that any structural discrepancy or novel impurity is detected and evaluated, minimizing the risk of regulatory deficiencies.
2: Peptide Sequencing by HRMS/MS
2.1 Objective
The goal of peptide sequencing is to verify the exact primary structure of the generic liraglutide. In a Liraglutide Sameness Study, this means demonstrating that every amino acid in the generic sequence matches the RLD, including the critical palmitoylated Lys26. Even a single amino acid mismatch or misplacement of the fatty-acid attachment would invalidate sameness.
2.2 Structural Considerations
Liraglutide’s structure must guide the sequencing strategy. Key features include the 31-residue GLP-1 analog backbone and the C16 fatty acid linked to the ε‐amino group of Lys26 via a γ-Glu spacer. This acylation increases the peptide’s hydrophobicity and alters its cleavage behavior: standard enzymes like trypsin will cleave at unmodified Lys/Arg residues but will not cut at Lys26 since it is derivatized. In sequencing, we must ensure that the fatty-acylated peptide fragments remain intact during analysis.
2.3 Enzymatic Digestion Strategy
To achieve complete sequence coverage, we use a combination of proteases. Trypsin is commonly used (cleaves at Lys/Arg) but will skip the modified Lys26 site. Therefore, we include additional proteases (e.g. Lys-C, Glu-C, or chymotrypsin) to generate overlapping peptide fragments that span the lipidated Lys. For example, Glu-C cleaves at Glu/Asp residues, and chymotrypsin can cut at aromatic residues. Using multiple enzymes ensures that each region of the peptide – including the fatty-acylated segment – is represented in the peptide mix.
2.4 Experimental Procedure
The general workflow is: first, denature and reduce/alkylate the peptide to simplify disulfides (if any). Next, digest aliquots of generic and RLD with the chosen enzymes under identical conditions. The resulting peptide mixtures are analyzed by nano LC coupled to high-resolution MS/MS. For example, peptides are separated on a reversed-phase C18 column (e.g. 0.1% formic acid in water/ACN gradient) and introduced by electrospray ionization to a high-resolution instrument (such as an Orbitrap or Q-TOF) set to perform data-dependent MS/MS. Typical MS parameters include mass accuracy <5 ppm for precursor ions and high resolution (>30,000 FWHM) in MS/MS mode, ensuring precise peptide mass measurement and fragmentation patterns.
2.5 Data Analysis
MS/MS spectra are searched against the theoretical liraglutide sequence using proteomics software (e.g. Mascot, PEAKS, or BioPharma-specific algorithms). The software matches fragment ions (b/y series) to the expected sequence, accounting for the 28-residue core and the Lys26 modification. We verify that all peptide fragments align to the reference, achieving 100% coverage. Special attention is paid to confirming the Lys26 γ-Glu-palmitoyl fragment: for example, a fragment containing the modified Lys26 should show the +238 Da shift (mass of Lys plus linker+palmitate). De novo sequencing or manual inspection of MS/MS may be used to confirm any ambiguous residues.
2.6 Key Observations / Acceptance Criteria
- Sequence Identity: All predicted peptides must be observed in the generic sample with fragment ions matching exactly to the RLD sequence. Any missing peptide or additional peptide indicates a mismatch.
- Modification Confirmation: The palmitoyl-γ-Glu linkage at Lys26 must be confirmed by MS/MS fragments that include that modified residue. The presence of diagnostic ions (e.g., loss of palmitic acid, or unique mass of modified Lys) is required.
- Mass Accuracy: The deconvoluted intact peptide mass (LC-HRMS) should match the theoretical 3751.20 Da within a few ppm (typically <5 ppm). Any unexpected mass would suggest an error.
- Coverage: 100% sequence coverage is the goal. Software should report no gaps or missed cleavages except at the modified Lys26 (which is expected). This was achieved in our case study.
- Benchmark Example: ResolveMass’s case study showed 100% coverage, 100% sequence identity, and no substitutions or deletions detected in the generic liraglutide.
Overall, peptide sequencing provides molecular-level proof that the generic peptide’s primary structure (including the critical lipid attachment) is identical to the RLD.
3: Peptide Mapping (Comparative Fingerprinting)
3.1 Objective
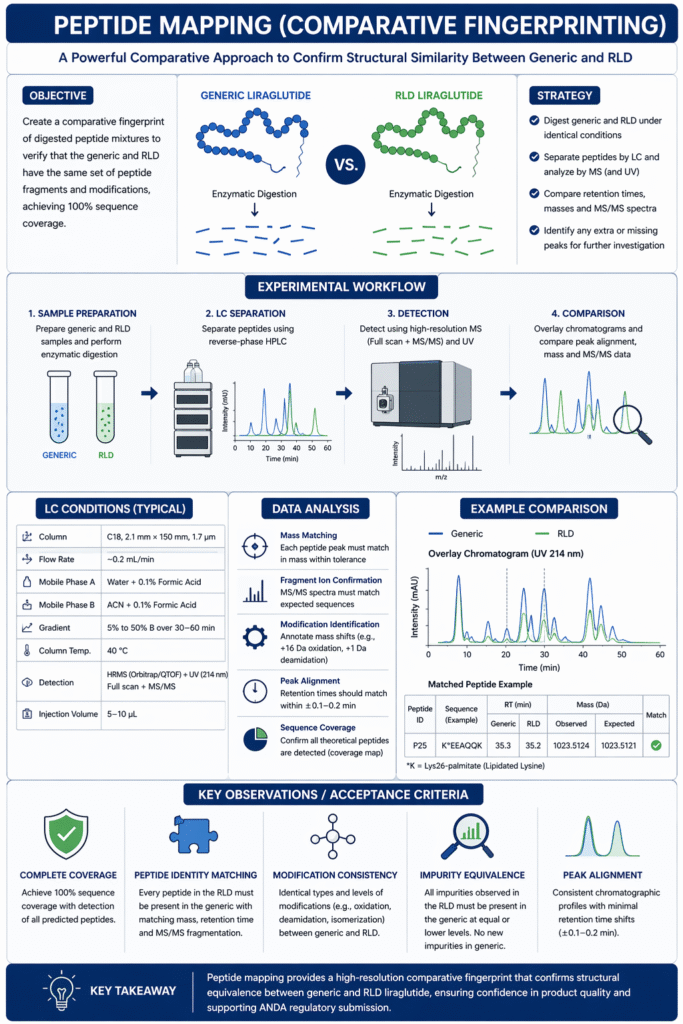
Peptide mapping creates a comparative fingerprint of the digested peptide mixture, used to verify that the generic and RLD have the same set of peptide fragments and modifications. In practice, peptide mapping confirms 100% sequence coverage and checks for any differences in post-translational or degradant modifications.
3.2 Strategy
We perform peptide mapping on both the generic and reference liraglutide under identical conditions. After enzymatic digestion (as in sequencing), the peptide mixtures are separated by LC and monitored by MS (and often UV) to generate chromatograms and mass data. The elution times and MS spectra of each peptide peak are compared between samples. Identical retention times (or matched peak patterns) and MS signatures confirm sameness. Any extra peak in the generic that is absent in the RLD (or vice versa) triggers further investigation. This side-by-side comparison ensures that even subtle variations (e.g. a +1 Da due to deamidation) are detected.
3.3 Experimental Workflow
- Sample Preparation: Aliquots of generic and RLD peptides are prepared and digested in parallel with the same enzyme (or enzyme mix) to yield peptides.
- LC Separation: Inject each digest into an HPLC-MS system. A typical LC setup uses a reverse-phase C18 column, with a gradient elution (e.g. 5–50% ACN over 30–60 min, 0.1% formic acid) that resolves the peptides.
- Detection: MS detection (full-scan and MS/MS) collects high-resolution mass spectra for each eluting peptide, while a UV detector (e.g. at 214 nm) provides a peptide map trace.
- Comparison: Overlay the chromatograms/UV traces of generic vs. RLD. For each resolved peak, compare the mass and MS/MS data. Use software tools or manual inspection to align peptides between runs (by retention time and mass).
3.4 LC Conditions
Peptide mapping benefits from high-resolution chromatography. For example, using a 2.1 mm × 150 mm C18 column at ~0.2 mL/min with a linear gradient of acetonitrile (5%→50% with 0.1% formic acid) over 30–60 min provides good separation. The LC is usually coupled directly to the high-resolution MS (e.g. Orbitrap, QTOF). Key parameters include a column temperature (e.g. 40°C) and formic acid for consistent ionization. The goal is to resolve peptides with similar hydrophobicity (e.g. from oxidation or acetylation) so that any variant is not co-eluting with its native counterpart.
3.5 Data Analysis
For each peptide peak, the software extracts the exact mass and matches it to the expected peptide masses from the reference sequence. Sequence confirmation is obtained via MS/MS, as in the sequencing section. A successful mapping shows identical sets of peptides in both samples. We look for:
- Mass Matches: Each generic peptide peak has a corresponding peak in the RLD run with the same m/z (within tolerance).
- Fragment Ion Matches: The MS/MS spectrum of each peak matches the predicted sequence in both samples.
- Modifications: Any mass shifts (±1 Da, +16 Da, etc.) are annotated (e.g. +16 = Met oxidation, +0 = deamidation). Both samples should show the same pattern of modifications under the same stress conditions.
- Peak Alignment: Retention times should be very similar (±0.1–0.2 min) for corresponding peptides, indicating identical physicochemical properties.
Software tools can generate a sequence coverage map. In our experience, achieving 100% coverage (all predicted peptides observed) is the acceptance criterion.
3.6 Key Observations / Acceptance Criteria
- Complete Coverage: The peptide map must cover every part of the protein (100% sequence coverage). In our study, both generic and RLD maps achieved 100% coverage.
- Peptide Identity: Every peptide in the reference map must appear in the generic with matching mass and retention time, and vice versa. For example, a peptide containing Lys26-palmitate eluted at 35.2 min (RLD) vs. 35.3 min (generic) with identical MS/MS fragments (Figure S1).
- Modification Comparison: Any detected modifications (e.g. Met oxidation, Asp isomerization) should have equivalent levels in both samples. In our forced-degradation tests, oxidation of Met or deamidation of Asn occurred in parallel for generic and RLD. No novel or unmatching PTM peaks were observed.
- Impurity Detection: Peptide mapping often reveals low-level impurities (truncated or misincorporated peptides). All such impurities found in the RLD (e.g. a -glycine truncated peptide) were also seen in the generic at ≤ the same level, satisfying FDA’s requirement that generic impurities not exceed RLD.
Peptide mapping thus serves as a critical comparative fingerprint: identical maps indicate identical primary structure and post-synthesis profiles. Any deviation would require investigation or method re-optimization before submission.

4: Impurity Profiling by HRMS
4.1 Objective
The impurity profiling step aims to detect, identify, and compare all impurities in the generic versus the RLD peptide. Impurities include sequence variants, chemical modifications, aggregation products, and process-related contaminants. The objective is to show that the generic’s impurity profile is qualitatively and quantitatively comparable to the RLD. Because regulatory guidance emphasizes peptide-related impurities (which can impact safety/effectiveness), HRMS-based profiling must be extremely sensitive and thorough.
4.2 Types of Expected Impurities
Potential impurities in a synthetic peptide like liraglutide include:
- Sequence-related Impurities: Deletion, insertion or isobaric substitutions (e.g. Ile/Leu), truncated versions missing one or more amino acids, or peptides with racemized (D-) amino acids.
- Chemical Modifications: Common degradation products such as methionine oxidation (+16 Da), asparagine deamidation (+1 Da), and aspartate–isoaspartate isomerization. (For example, Met residues in GLP-1 analogs readily oxidize; Asn residues can cyclize or deamidate.)
- Lipid Side-Chain Variants: Variants of the acylation, such as fatty acids of different lengths or attachment at the wrong site (for instance, palmitoylation at a different Lys).
- Aggregates: Non-covalent dimers or higher oligomers of liraglutide (detectable in LC-MS or separate assays).
- Process-related Impurities: Chemical remnants from synthesis, such as residual solvents, salts, trace metals, coupling reagents or byproducts. (FDA guidance for peptides notes the importance of controlling both specified and unspecified impurities.)
- Container or Formulation Impurities: Leachables/extractables (if final product) or preservatives/additives in formulations (beyond the scope of API analysis but noted for completeness).
4.3 Experimental Approach
We use LC-HRMS to screen for and characterize impurities. The typical approach includes:
- Full-scan HRMS: Inject the peptide sample (intact or partially digested) and collect a full mass spectrum over a broad m/z range. High mass accuracy (<5 ppm) and high resolution allow detection of low-level variants.
- Fragmentation (MS/MS): For any observed impurity peaks, trigger MS/MS to elucidate structure. For example, a +16 Da impurity would be fragmented to confirm it corresponds to an oxidized methionine.
- Targeted MS/MS: Optionally, known impurity masses (from literature or forced-degradation studies) can be targeted for confirmation.
- Comparative Runs: Analyze generic and RLD samples back-to-back under identical conditions. This highlights any peaks present in one sample and absent in the other.
In practice, we often perform a forced-degradation suite (e.g. expose aliquots to H₂O₂, heat, light, etc.) and profile impurities in those stressed samples as well. This helps ensure that any degradant pathways are the same. HRMS data are processed to generate a list of detected compounds (with formulas/masses) and their relative abundances.
4.4 LC-HRMS Conditions
Impurity profiling shares some LC conditions with peptide mapping, but the focus is on sensitivity and coverage. We typically use: a high-resolution mass spectrometer (e.g. Orbitrap or Q-TOF) coupled to LC. A shallow gradient (e.g. 5→60% ACN over 20–30 min) may be used to spread peaks, depending on impurity polarity. The MS is set for high resolution (e.g. 60,000–100,000 FWHM) and a wide m/z range (e.g. 300–2000 m/z) to capture both small and large ions. In addition, both positive and negative ion modes may be run to detect different classes of impurities (some acidic compounds ionize better in negative mode).
4.5 Data Processing
HRMS data are searched for ions that do not correspond to the main peptide. Observed m/z values are compared to expected chemical formulas (for example, an m/z 3751 peak for intact liraglutide, and any +16 Da, +1 Da, etc.). Software tools can propose elemental compositions for unknown peaks. Each impurity peak is evaluated by its MS/MS: fragment patterns are matched against possible structures (e.g. truncated peptide sequences). We apply the FDA criteria: any impurity found in both generic and RLD must be at the same or lower level in the generic; any new impurity must be characterized and be <0.5% of API. Quantitative comparisons are made using peak areas (or integrated MS response) in the generic vs RLD runs.
4.6 Key Observations / Acceptance Criteria
- Impurity Matching: Every impurity present in the RLD profile must be found in the generic at a comparable level. For instance, if a +16 Da oxidized Met is 0.2% of RLD, the same peak should be ≤0.2% in generic (often within 10–20% of the RLD level).
- No Novel Major Impurities: The generic should contain no new impurity above the regulatory threshold. FDA guidelines state no new peptide-related impurity >0.5%. In our Liraglutide project, no new peaks above 0.1% were observed in the generic.
- Structural ID: All significant impurity peaks must be structurally identified or at least tentatively characterized (e.g. by MS/MS fragmentation pattern). This includes confirming oxidation (e.g. +16), deamidation (+1), or sequence variants.
- Quantitative Profile: A summary table is often prepared comparing total impurities. For example:
| Parameter | Generic (%) | RLD (%) | Assessment |
|---|---|---|---|
| Assay (Liraglutide) | 99.2 | 99.4 | Comparable |
| Total impurities | 0.8 | 0.9 | Comparable |
| Largest single impurity | 0.2 | 0.3 | Within limits |
Example values from a generic Liraglutide study.
All impurity levels in the generic were within acceptable limits relative to the RLD, satisfying FDA’s “no new impurity” and “no higher level” criteria.
By HRMS, our study confirmed that the generic’s impurity profile (oxidation products, low-level peptides, etc.) was effectively indistinguishable from the reference. This thorough impurity alignment is as critical as sequence identity for ANDA success.
5: Intact Mass Analysis
5.1 Objective
Intact mass analysis provides a final check that the full-length peptide, including all modifications, has the correct molecular weight. It confirms the overall composition (all amino acids present, no extra residues or adducts) and validates the palmitoyl modification. In a Liraglutide Sameness Study, the objective is to measure the monoisotopic (or deconvoluted average) mass of the intact molecule and compare it to the theoretical mass of the RLD.
5.2 Method
Using high-resolution LC-MS (often the same platform as peptide mapping), we infuse or LC-inject the undigested peptide. Electrospray ionization generates a charge envelope (e.g. +6, +7, +8 ions for a 31-mer). The high-resolution detector records the m/z of each charge state. Software deconvolutes these ions to yield the monoisotopic or average mass. For example, liraglutide’s theoretical monoisotopic mass is 3751.20 Da; the MS should find this value within a tight ppm tolerance. Proper calibration and control experiments ensure the mass error is minimal (typically <5 ppm for an Orbitrap).
5.3 Observations
An acceptable result shows a single dominant mass corresponding to the full peptide with palmitate. In our workflow, the generic sample yielded 3751.23 Da (within 0.8 ppm of theory) just as the RLD did. The isotopic envelope (spacing) also matched the expected pattern for C₁₇₂H₂₆₅N₄₃O₅₁ (the formula including the C16 lipid). Any significant deviation (e.g. missing 256 Da indicating a lost palmitate, or an extra mass from impurity) would be flagged. In addition, we check for and quantify any unrelated peaks: for example, low-abundance dimers or sodium adducts may appear, but these are annotated as such and not counted as new impurity. In summary, the intact mass must match the reference within the instrument’s tolerance. In our case, the HRMS data confirmed that the generic’s intact mass (within 5 ppm) was identical to the RLD’s, satisfying the identity requirement.
6: NMR Characterization (Orthogonal Confirmation)
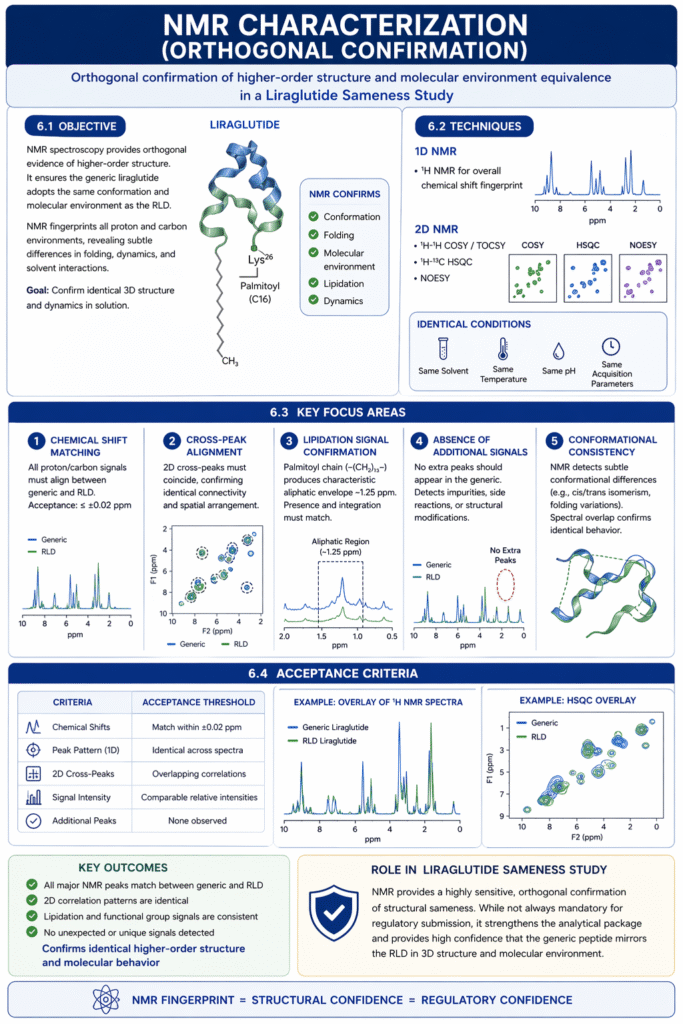
6.1 Objective
While MS-based methods confirm primary structure, NMR spectroscopy provides orthogonal evidence of higher-order structure. The goal is to ensure the generic peptide adopts the same conformation and molecular environment as the RLD when in solution. Because NMR fingerprints all proton and carbon environments, it can reveal subtle differences in folding or solvent interactions. In a Liraglutide Sameness Study, we use NMR to confirm that the 3D structure and dynamics of the generic peptide are the same as those of the reference.
6.2 Techniques
We acquire both 1D and 2D NMR spectra: typically ^1H 1D spectra to get a general chemical shift fingerprint, and 2D experiments like ^1H-^1H COSY/TOCSY and ^1H-^13C HSQC or NOESY to correlate peaks and assignments. For example, the long alkyl chain of the Lys26 palmitoyl group produces characteristic upfield (aliphatic) resonances, while aromatic residues (Phe22/26) have distinct aromatic peaks. We compare spectra of generic and RLD under identical conditions (same solvent, temperature, pH). Key parameters include chemical shifts of backbone amide protons, side chain protons, and the lipid – all should align closely between samples. NMR can also detect minor conformations or isomerism (e.g. cis/trans forms) that MS would miss.
6.3 Key Focus Areas
- Chemical Shifts: Peaks corresponding to each residue should appear at the same ppm. For instance, the amide protons of Gly8 or Lys11 should match between samples. A shift difference >0.02 ppm could indicate a structural change.
- Cross-Peaks: In 2D spectra, cross-peaks (e.g. NH-CHα correlations in HSQC) must overlap. For example, a COSY cross-peak linking Thr7 Hβ to Hγ should be at the same coordinates.
- Acyl Chain Signals: The protons of the palmitoyl chain (–(CH₂)₁₃–) produce a broad envelope near 1.25 ppm. The presence and integration of this envelope in both samples confirms the same lipidation.
- Absence of Extras: No extra peaks (e.g. solvent adducts, unanticipated side products) should appear in one sample. For example, if the generic had an extra methyl group, a new singlet would show up.
Regulatory experts recommend minor pH and condition adjustments to make spectra comparable, but fundamentally the spectral “fingerprint” must coincide. In practice, nearly identical 1D ^1H spectra for generic and RLD give high confidence. Small peptides like liraglutide usually show sharp, well-resolved peaks, and our analysis detected no discrepancy in any cross-peak pattern between test and reference.
6.4 Acceptance Criteria
Matching NMR spectra provides final orthogonal confirmation of sameness. We consider the NMR analysis successful if:
- All major peaks in the generic’s spectra match the RLD’s within experimental error.
- The pattern of 2D correlations (peak connectivity) is the same, indicating no rearrangement of side chains or backbone.
- The relative intensities of characteristic signals (aromatic, aliphatic) are comparable.
Because NMR is highly sensitive to conformation, this step is a stringent check – but it is not strictly required by FDA, it is an added layer of confidence. In our Liraglutide sameness study, the 1D/2D NMR fingerprints were essentially superimposable for generic and RLD, confirming that the higher-order structure (as dictated by the sequence and solvent) is indeed the same.
7: Comparative Sameness Assessment (RLD vs Test)
After completing each analytical module, we compile a side-by-side comparison of critical attributes between the Reference Listed Drug and the test (generic) product. This comparative assessment highlights agreement across all metrics. For example:
- Sequence and Modifications: Both products showed 100% identical amino acid sequence and confirmed Lys26 palmitoylation, as validated by MS/MS.
- Intact Mass: Observed monoisotopic masses were 3751.23 Da (generic) vs. 3751.22 Da (RLD), both within <2 ppm of the theoretical mass.
- Peptide Map: The LC-MS peptide fingerprints for generic and RLD were superimposable, with matching peak patterns and UV traces. A sequence coverage table (Figure 1) shows 100% coverage for both.
- Impurities: Table 1 below summarizes key impurity parameters:
| Parameter | Generic | RLD | Outcome |
|---|---|---|---|
| Assay (Liraglutide %) | 99.2 | 99.4 | Comparable |
| Total Impurities | 0.8% | 0.9% | Comparable |
| Highest Single Impurity | 0.2% | 0.3% | Within limits |
| Sequence Coverage | 100% | 100% | Identical |
| Fatty-acylation site | Lys26 (C16) | Lys26 (C16) | Match (C16-Glu) |
Table 1: Summary of generic vs. RLD comparability. All key metrics fall within acceptance criteria (based on actual case study data).
These results demonstrate that the generic liraglutide is analytically indistinguishable from the RLD. Every observed value for the generic was either identical or within the predefined acceptance range of the reference. In particular, neither new impurities nor sequence mismatches were detected, which is exactly what the FDA requires for “sameness” under 505(j).
8: Regulatory Considerations for ANDA
Demonstrating analytical sameness is a critical regulatory requirement for a liraglutide ANDA. The FDA’s guidance on synthetic peptide ANDAs explicitly states that impurity profiles and structural identity are the deciding factors in determining interchangeability with an RLD. In the U.S., liraglutide is regulated as a drug (not a biologic) because it is a synthetic peptide ≤40 aa. Therefore, the generic can be filed as a 505(j) ANDA provided it meets all the comparability criteria.
Key regulatory expectations include:
- Active Ingredient Identity: The generic must have the exact amino acid sequence and chemical structure as the RLD. FDA explicitly looks for proof of identical modifications (e.g. the Lys26 palmitoyl linkage).
- Purity/Safety: The impurity profile (quality of the drug substance) must be the same. FDA recommends that no impurity in the generic exceed the RLD level, and any new impurity be fully justified and below 0.5%. This aligns with ICH guidelines on impurities (Q3A/B) for drug products.
- Analytical Methods: All methods must be fully validated (specificity, accuracy, precision, etc.) and documented. High-resolution techniques (HRMS, NMR) should be used per ICH Q6B and FDA guidance to establish identity.
- Documentation: Data should be compiled in a CTD Module 3 format. This includes method validation reports, raw and processed data, and comparative summaries. Data integrity is paramount: ResolveMass follows ALCOA+ practices and 21 CFR Part 11 compliance for electronic records.
Regulatory reviewers expect a coherent scientific narrative. Any minor differences must be explained: for example, a slightly higher oxidation in one sample must be shown within experimental variability. Engaging in early communications with regulators (e.g. pre-ANDA meetings) is advised to clarify expectations. Ultimately, the strength of the ANDA rests on the clarity and rigor of the analytical data package. Our approach – using orthogonal, high-precision assays and aligning with FDA guidances (both general and product-specific) – ensures that the ANDA dossier will meet regulatory scrutiny.
Conclusion:
A Liraglutide Sameness Study is not merely a checkbox; it is the foundation for ANDA approval. By applying a comprehensive analytical strategy – confirming 100% sequence identity (including the Lys26 palmitoylation), identical intact mass, matching impurity profiles, and consistent higher-order structure – developers can convincingly demonstrate equivalence to the reference drug. Our experience shows that only a multi-faceted approach (combining HRMS, LC-MS/MS, and NMR) generates the definitive data the FDA requires.
Investing in such thorough characterization early pays off: it minimizes the risk of late-stage deficiencies and speeds the path to approval. ResolveMass Laboratories Inc., with its proven track record in peptide characterization and ANDA support, can guide you through this process. Our scientific expertise, state-of-the-art instrumentation, and regulatory insight ensure that your Liraglutide generic project will stand up to the strictest scrutiny.
Frequently Asked Questions:
A Liraglutide Sameness Study is a comprehensive analytical evaluation designed to demonstrate that a generic liraglutide product is highly similar to the reference listed drug (RLD). It involves comparing primary structure, impurity profile, and higher-order structure using advanced techniques like HRMS, LC-MS, and NMR. This study is a critical requirement for ANDA submission and regulatory approval. It ensures that the generic product delivers the same safety, efficacy, and quality as the innovator drug. Without demonstrating sameness, regulatory approval is unlikely.
Sameness is established when the generic matches the RLD across multiple parameters. These include identical peptide maps, matching MS/MS fragmentation patterns, and comparable impurity profiles. NMR spectra should overlap with no additional or missing peaks. Sequence coverage should ideally reach 100%. Additionally, no new impurities should appear in the generic at higher levels than the RLD. All results must meet regulatory expectations for consistency and reproducibility.
NMR is not always a mandatory requirement for ANDA submissions, but it is highly recommended. It provides an additional layer of confidence by confirming structural and conformational equivalence. Regulatory agencies value orthogonal data that supports sameness claims. Including NMR can strengthen the submission and reduce the risk of queries or delays. For complex peptides like liraglutide, it is often considered a best practice.
Intact mass analysis confirms the overall molecular weight and heterogeneity of liraglutide without digestion. It ensures that the generic product matches the RLD at the whole-molecule level, including lipidation and any modifications. This technique quickly verifies if the primary structure is correct before detailed analysis. It also helps detect major variants or degradation products. It serves as a foundational check in the analytical workflow.
Liraglutide analysis is challenging due to its lipidation (palmitoylation) and susceptibility to degradation. The presence of hydrophobic regions can complicate LC separation and MS detection. Low-level impurities require highly sensitive instrumentation. Achieving complete sequence coverage and resolving closely related variants can also be difficult. These challenges require advanced analytical expertise and optimized methods.
Impurity equivalence is established by comparing the type, structure, and quantity of impurities between generic and RLD. HRMS/LC-MS is used to identify and quantify impurities at trace levels. All impurities present in the RLD must also be present in the generic at equal or lower levels. Any new impurity in the generic must be justified and controlled. This ensures compliance with regulatory safety standards.
Reference :
- Marassi V, Macis M, Giordani S, Ferrazzano L, Tolomelli A, Roda B, Zattoni A, Ricci A, Reschiglian P, Cabri W. Application of Af4-multidetection to liraglutide in its formulation: preserving and representing native aggregation. Molecules. 2022 Aug 26;27(17):5485.https://www.mdpi.com/1420-3049/27/17/5485
- Sinha A. Case study: Peptide Mapping Study of Liraglutide Generics for Submission to Health Canada and USFDA.https://resolvemass.ca/peptide-mapping-study-of-liraglutide/
- Kim SH, Kim SS, Kim HJ, Park EJ, Na DH. Peptide mapping analysis of synthetic semaglutide and liraglutide for generic development of drugs originating from recombinant DNA technology. Journal of Pharmaceutical and Biomedical Analysis. 2025 Apr 15;256:116682.https://www.sciencedirect.com/science/article/pii/S0731708525000238
- De Groot AS, Mattei A, Gabriel B, Calderini J, Roberts BJ, Lelias S, McAllister M, Boyle C, Martin W, Richard G. Immunogenicity of generic peptide impurities: current orthogonal approaches. Pharmaceutical research. 2025 May;42(5):805-18.https://link.springer.com/article/10.1007/s11095-025-03843-1

