Introduction: The Analytical Burden of Sequence Confirmation in Biosimilars
Peptide Mapping in Biosimilars plays a critical role during development, but its purpose here is different from early discovery work. In biosimilars, the amino acid sequence is already known from the reference product. The main goal is not to discover the sequence, but to confirm that it has been copied exactly and to measure how similar it is.
Because of this, peptide mapping becomes a comparison-based analytical task. Each peptide that is detected must align with the reference sequence. Every modification site needs to be measured and compared, and any unexpected mass signal must be carefully reviewed as a possible sequence variant.
There is very little room for error. Regulatory agencies expect full sequence confirmation. Partial sequence coverage is not acceptable for proving similarity. In addition, any identified variant must be fully studied, including its structural impact and any possible risk of immunogenicity.
Explore our specialized services: Expert Biosimilar Characterization Using Mass Spectrometry
Share via:
Article Summary
- Peptide Mapping in Biosimilars is not just a regulatory step—it is a high-resolution analytical approach used to confirm primary sequence identity and detect low-level differences from the reference product.
- LC-MS/MS-based peptide mapping should achieve at least 95% sequence coverage. Orthogonal digestion strategies (e.g., trypsin with Lys-C or Glu-C) are used to address missed cleavages and overlapping peptides.
- Sequence variants such as amino acid substitutions, misincorporation events, and C-terminal lysine clipping must be clearly distinguished from analytical artifacts, often using intact mass correlation.
- Post-translational modification (PTM) analysis goes beyond common changes like deamidation and oxidation. It includes site-specific glycoform profiling, N-terminal pyroglutamate quantification, and evaluation of disulfide bond integrity.
- Regulatory bodies such as the U.S. Food and Drug Administration, European Medicines Agency, and Health Canada require peptide mapping as part of biosimilar comparability, with scientifically justified analytical strategies.
- Data analysis plays a critical role, where bioinformatics tools—including spectral matching algorithms and false discovery rate (FDR) controls—ensure reliable variant identification.
- ResolveMass Laboratories Inc. provides validated, GMP-aligned peptide mapping workflows tailored for complex biologics such as monoclonal antibodies and Fc-fusion proteins.
Why Standard Tryptic Digestion Is Insufficient for Peptide Mapping in Biosimilars
Using only trypsin digestion often results in incomplete sequence coverage. In Peptide Mapping in Biosimilars, this is not a small issue but a serious regulatory concern. Some regions do not digest well with trypsin, such as areas near proline residues, highly hydrophobic regions, and sections with strong disulfide bonding. These regions often produce large peptides that ionize poorly and are difficult to detect.
More reliable strategies include:
Multi-Enzyme Approaches in Peptide Mapping in Biosimilars
Combining trypsin with enzymes like Lys-C under denaturing conditions improves digestion efficiency. This helps reduce missed cleavages, especially in areas with glycosylated asparagine residues.
Orthogonal Digestion Methods for Better Coverage
Using enzymes such as Glu-C, Asp-N, and chymotrypsin produces different peptide fragments. Running these digestions together and comparing the peptide maps helps fill sequence gaps.
Optimized Reduction and Alkylation
If disulfide bonds are not fully reduced, peptide recovery is affected. The choice of alkylating agents, such as iodoacetamide or N-ethylmaleimide, creates specific mass changes that must be considered during analysis to avoid incorrect conclusions.
| Enzyme | Primary Cleavage Site | Key Application in Biosimilars |
|---|---|---|
| Trypsin | C-terminal Arg, Lys | Backbone coverage, CDR peptides |
| Lys-C | C-terminal Lys only | Resolves Arg-Pro missed cleavages |
| Glu-C | C-terminal Glu (pH 4) or Glu/Asp (pH 8) | Fc region coverage, N-terminal variants |
| Asp-N | N-terminal Asp | Complementary to trypsin for challenging sequences |
| Chymotrypsin | Phe, Tyr, Trp, Leu | Hydrophobic domain coverage |
There is no single digestion method that works for every biosimilar. Each Peptide Mapping in Biosimilars workflow must be adjusted based on the molecule’s structure.
Learn how we confirm similarity: Prove Biosimilarity Using Advanced LC-MS Techniques
Sequence Variant Detection: Separating True Variants from Analytical Noise
Detecting sequence variants is one of the most important parts of peptide mapping. Even one amino acid change can impact binding, stability, or immune response.
The real challenge is not finding mass differences, but deciding whether they are true variants or just analytical errors.
Common sequence variants include:
Amino Acid Substitutions
These appear as changes in peptide mass. Some substitutions have nearly identical masses, so high-resolution MS/MS is required for correct identification.
Misincorporation Events
These are usually present at very low levels, often below 1%. Careful analysis using extracted ion chromatograms and signal-to-noise checks is needed to confirm them.
C-Terminal Lysine Clipping
This is often seen in monoclonal antibodies produced in CHO cells. It must be measured and compared with the reference product.
N-Terminal Processing Variants
Incomplete signal peptide removal can be verified by studying the N-terminal peptide region.
An important validation step is comparing peptide-level data with intact protein mass results. If a detected variant does not show a matching shift in intact mass based on its level, it is likely not a true variant.
False discovery rates must be strictly controlled, usually at or below 1%, using methods like target-decoy strategies. At very low levels, incorrect matches become more likely, so strong statistical control is necessary.
Need high-level protein analysis? Comprehensive Intact Mass Analysis for Biosimilars
PTM Site Localization in Peptide Mapping in Biosimilars: Beyond Deamidation and Oxidation
For biosimilar comparison, analyzing post-translational modifications (PTMs) is not just about detecting them. It is about confirming that their type, location, and quantity match the reference product within acceptable limits.
Key PTMs that require detailed study include:
Deamidation
This is especially important in functional regions like complementarity-determining regions (CDRs). Accurate localization requires MS/MS data to identify the exact residue.
Oxidation
Methionine oxidation, particularly in the Fc region, can affect binding and half-life. Site-specific measurement is required.
N-Terminal Pyroglutamate Formation
This occurs when glutamine cyclizes. If present in the reference product, it must also be measured in the biosimilar.
Glycation
This non-enzymatic modification adds sugar molecules to lysine residues, creating measurable mass shifts and increasing heterogeneity.
Disulfide Bond Scrambling
Incorrect pairing of disulfide bonds creates new peptide forms. These are best detected by comparing reduced and non-reduced peptide maps.
Advanced support for complex molecules: Generic Peptide & Oligonucleotide Projects
Glycopeptide Analysis Within the Peptide Map: Site-Specific Glycoform Profiling
Glycosylation is a key quality attribute in many biologics, especially monoclonal antibodies. Studying glycopeptides provides insights that cannot be gained from released glycan analysis alone.
Key benefits of glycopeptide analysis include:
- Confirmation that glycosylation occurs at the correct site
- Detection of non-glycosylated forms along with glycosylated ones
- Identification of differences in site occupancy when multiple glycosylation sites exist
Accurate analysis requires advanced fragmentation techniques such as data-independent acquisition or stepped collision energy. These methods produce both glycan and peptide fragment ions in a single run, allowing full characterization.
Targeted peptide development: Specialized Generic Peptide Semaglutide Projects and Liraglutide Generic Development Services
Regulatory Expectations for Peptide Mapping in Biosimilars
Regulatory agencies focus more on results than on specific methods. The expectation is clear: complete and scientifically supported sequence confirmation, along with quantitative data on variants and PTMs.
Typical expectations include:
| Regulatory Requirement | Specific Expectation |
|---|---|
| Sequence coverage | ≥95% of theoretical protein sequence confirmed |
| Variant detection limit | Sensitivity down to 0.1% relative abundance for substitutions |
| PTM comparability | Site-specific quantification with predefined acceptance criteria |
| Method validation | ICH Q2(R2)-aligned validation parameters documented |
| Orthogonal data | Intact mass correlation required to support peptide-level findings |
| Reference standard | Characterized reference product lots used as comparator in the same analytical run |
A common issue in submissions is not defining detection limits. Simply stating that no variants were found is not enough without showing the sensitivity of the method used.
Navigate the pathway to approval: Regulatory Support for Generic Drug Development and Lanreotide Generic Development Services
Bioinformatic Rigor in Peptide Mapping in Biosimilars
The accuracy of peptide mapping depends heavily on data analysis. Modern mass spectrometry produces large datasets that require automated tools, but these tools must be carefully validated.
Important factors include:
Database Design
The database must include the exact reference sequence along with all expected modifications and possible variants.
Spectral Libraries
Using in-house spectral libraries increases confidence in identifying peptides, especially those present at low levels.
Chromatographic Alignment
Proper alignment of retention times ensures that observed differences are real and not due to instrument variation.
Isotope Pattern Analysis
Checking isotope distributions adds another level of confidence, especially for large or highly charged peptides.
Ensure analytical precision: Analytical Services for Generic Drug Development
Conclusion: Peptide Mapping in Biosimilars as a Strategic Comparability Tool
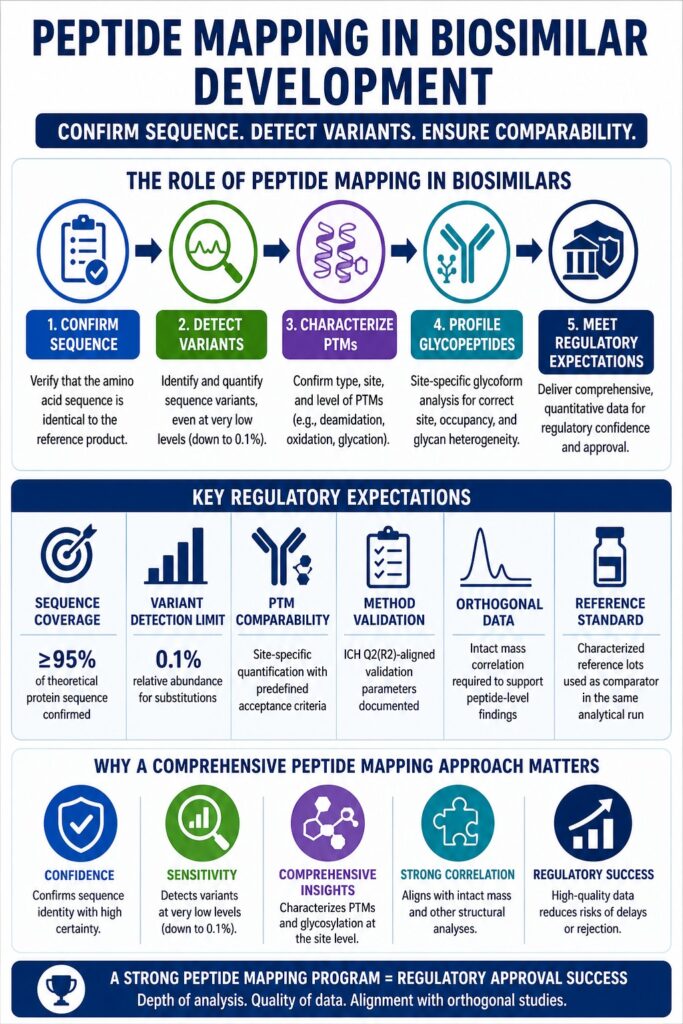
Peptide Mapping in Biosimilars is more than just a routine test. It provides detailed structural information at a very high level of resolution. When done correctly, it confirms sequence identity, measures variants, identifies modifications, and connects glycosylation patterns to specific sites.
The success of a peptide mapping strategy depends on three key elements: the depth of analysis, the quality of quantitative data, and how well the results align with other structural studies such as intact mass and higher-order structure analysis.
A strong peptide mapping program supports regulatory approval, while gaps in data often lead to delays or rejection. Success requires careful planning, optimized methods, and reliable data interpretation at every step.
Scale your project for success: Formulation Development and Manufacturing Scale-Up for Generic Drugs
📩 Ready to discuss your biosimilar peptide mapping requirements?
Contact ResolveMass Laboratories Inc. →
Frequently Asked Questions (FAQs) on Peptide Mapping in Biosimilars
In Peptide Mapping in Biosimilars, regulatory bodies such as the FDA and EMA generally expect at least 95% sequence coverage, while the ideal scientific goal is full coverage. Achieving this level often requires combining multiple enzymes, as a single digestion method is usually not enough for complex biologics. Any missing regions must be clearly explained with proper justification. If gaps are not addressed, they may raise concerns during regulatory review.
Isobaric amino acids, such as leucine and isoleucine, are difficult to differentiate because they share the same mass. In Peptide Mapping in Biosimilars, advanced techniques like Electron Transfer Dissociation (ETD) or high-resolution mass spectrometry are used to separate these closely related signals. In some cases, known genetic sequence data is also used to support identification. Careful data analysis is essential to avoid incorrect assignments.
C-terminal lysine clipping is a typical modification seen in many monoclonal antibodies, especially those produced in CHO cells. In Peptide Mapping in Biosimilars, its importance lies in comparing how much clipping occurs in both the biosimilar and the reference product. Differences in levels may affect charge profiles and, in some cases, biological interactions. Therefore, it must be measured and evaluated carefully.
Peptide mapping is an essential tool, but it cannot confirm biosimilarity on its own. In Peptide Mapping in Biosimilars, it mainly provides detailed information about primary structure and modifications. A full comparability assessment also requires additional studies, such as intact mass analysis, higher-order structure evaluation, and functional testing. These combined results provide a complete understanding of similarity.
Detecting very low-level variants requires sensitive analytical methods and strong data processing. In Peptide Mapping in Biosimilars, high-performance mass spectrometers and optimized sample preparation help capture signals below 0.5% abundance. Data must be validated using chromatograms, isotope patterns, and fragmentation results. This ensures that detected variants are real and not due to background noise.
Reducing peptide mapping involves breaking disulfide bonds before digestion, which helps confirm the main amino acid sequence. Non-reducing peptide mapping keeps these bonds intact, allowing analysis of disulfide linkages. In Peptide Mapping in Biosimilars, both approaches are important—one for sequence confirmation and the other for checking correct structural bonding. Together, they provide a more complete structural profile.
Released glycan analysis shows the overall glycan composition but does not indicate where each glycan is attached. In Peptide Mapping in Biosimilars, glycopeptide analysis connects each glycan to its exact site on the protein. This helps detect unexpected glycosylation and differences in site occupancy. It is especially important for proteins with multiple glycosylation sites.
Reference:
- Segu, Z., Stone, T., Berdugo, C., Roberts, A., Doud, E., & Li, Y. (2020). A rapid method for relative quantification of N-glycans from a therapeutic monoclonal antibody during trastuzumab biosimilar development. MAbs, 12(1), 1750794. https://pmc.ncbi.nlm.nih.gov/articles/PMC7188402/
- Nupur, N., Joshi, S., Guillarme, D., & Rathore, A. S. (2022). Analytical similarity assessment of biosimilars: Global regulatory landscape, recent studies and major advancements in orthogonal platforms. Frontiers in Bioengineering and Biotechnology, 10, 832059. https://pmc.ncbi.nlm.nih.gov/articles/PMC8865741/
- D’Atri, V., Guillarme, D., & Beck, A. (2025). Biopharmaceutical analysis—current analytical challenges, limitations, and perspectives. Analytical and Bioanalytical Chemistry. https://pmc.ncbi.nlm.nih.gov/articles/PMC12783247/



