Summary:
A Tirzepatide Sameness Study is the scientific backbone of a successful generic Abbreviated New Drug Application (ANDA). This report outlines how ResolveMass Laboratories Inc. utilizes orthogonal high-resolution techniques to prove structural identity between a generic synthetic peptide and the reference drug.
- Definition: Sameness is established by proving identical primary sequence, physicochemical properties, and higher-order structure (HOS) compared to the Reference Listed Drug (RLD).
- Regulatory Standard: Success depends on identifying all peptide-related impurities 0.10% and ensuring new impurities do not exceed 0.5%.
- Analytical Core: The study employs LC-HRMS for sequencing, multi-enzymatic mapping for fingerprinting, and 1D/2D NMR for atomic-level conformation proof.
- ResolveMass Edge: Our ISO 9001:2015 certified Canadian laboratory provides PhD-led expertise and state-of-the-art instrumentation (e.g., Orbitrap HRMS, 500 MHz NMR) to de-risk generic submissions.
1. Introduction
A Tirzepatide Sameness Study is a comprehensive analytical program designed to demonstrate that a generic synthetic version of Tirzepatide is molecularly identical to the innovator’s reference listed drug (RLD). This process is a critical component of FDA-aligned submissions and builds upon established frameworks such as Sameness Evaluation in ANDA and Active Ingredient Sameness in ANDA.
For complex peptides, manufacturers must follow FDA Peptide Sameness Study Requirements and adopt Analytical Strategies for Sameness Study to ensure regulatory success.
Tirzepatide, a 39-amino acid dual agonist, presents unique analytical challenges similar to other GLP-1 analogs such as Liraglutide Sameness Study, Semaglutide Sameness Study, and Exenatide Sameness Study.
2. Analytical Strategy Overview (Multi-Technique Approach)
The analytical strategy for a Tirzepatide Sameness Study relies on a multi-layered, orthogonal approach that combines high-resolution mass spectrometry (HRMS) with multidimensional NMR to eliminate any analytical blind spots. This strategy ensures that the primary sequence, impurity profile, and higher-order structure (HOS) are characterized using independent physical principles, as recommended by global regulatory agencies.
A Tirzepatide Sameness Study relies on orthogonal analytical techniques to eliminate blind spots. This aligns with best practices described in Orthogonal Analytical Techniques for ANDA Sameness Evaluation.
| Analysis Type | Primary Techniques | Target Attributes |
| Primary Structure | LC-HRMS/MS, Multi-Enzymatic Mapping | Amino acid sequence, site-specific acylation |
| Impurity Profile | LC-HRMS (Orbitrap/Q-TOF), MAM | Deletions, insertions, sequence variants 0.10% |
| Higher-Order Structure | 1D/2D NMR (HSQC, TOCSY, NOESY) | Secondary/tertiary folding, conformational sameness |
| Physicochemical | DSC, XRPD, SEC-MALS | Aggregation state, solubility, polymorphic form |
This structured approach is widely used in Sameness Evaluation of Synthetic Peptides for ANDA.
3. Peptide Sequencing by HRMS/MS
Peptide sequencing confirms the exact amino acid sequence and verifies structural identity with the RLD.
The approach follows similar principles used in Octreotide Sameness Study for ANDA Submission and Liraglutide Sameness Study for ANDA Submission.
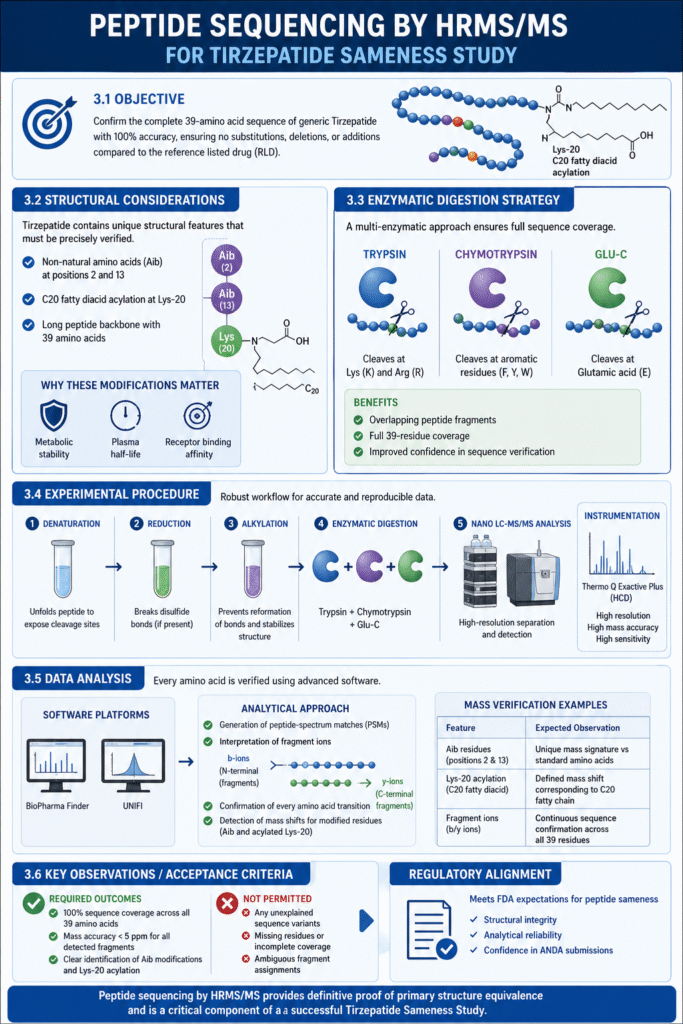
3.1 Objective
The primary objective is to confirm the 39-amino acid sequence of the generic Tirzepatide with 100% accuracy, ensuring no substitutions, deletions, or additions exist relative to the RLD.
3.2 Structural Considerations
Tirzepatide’s sequence includes non-natural amino acids (Aib) at positions 2 and 13 and a complex C_{20} fatty diacid acylation at Lys-20. These modifications are critical for metabolic stability and must be verified as correctly positioned and chemically intact during the Tirzepatide Sameness Study.
3.3 Enzymatic Digestion Strategy
To achieve full sequence coverage, ResolveMass employs a multi-enzymatic approach using Trypsin, Chymotrypsin, and Glu-C. This is necessary because Tirzepatide’s acylation can sterically hinder certain cleavage sites, and multiple enzymes provide overlapping fragments that verify the entire 39-residue span.
3.4 Experimental Procedure
Samples undergo denaturation, reduction, and alkylation before digestion. The resulting peptides are analyzed via nanoLC-MS/MS using high-energy collisional dissociation (HCD) on systems like the Thermo Q Exactive Plus.
3.5 Data Analysis
Sequencing data is processed using software such as BioPharma Finder or UNIFI, which generates a peptide-spectrum map. We analyze b-ions and y-ions to verify every amino acid transition, specifically confirming the mass shift associated with the acylated Lysine-20.
3.6 Key Observations / Acceptance Criteria
The sequence must show 100% coverage with a mass accuracy of < 5 ppm for all fragments. No unexplained sequence variants or missing residues are permitted under FDA sameness criteria.
4. Peptide Mapping (Comparative Fingerprinting)
Peptide mapping creates a molecular fingerprint proving identical fragmentation patterns.
This approach aligns with advanced workflows used in Peptide Sameness Study for ANDA.
4.1 Objective
The objective is to create a high-resolution “molecular fingerprint” that proves the generic product and RLD possess identical fragment patterns when subjected to identical enzymatic stress.
4.2 Strategy
We utilize a Comparative Multi-Attribute Method (MAM) where UV and MS chromatograms are overlaid. This allows for the simultaneous assessment of identity and relative abundance of all peptide components.
4.3 Experimental Workflow
Identical batches of generic Tirzepatide and RLD are digested in parallel under controlled conditions. The digests are injected into a UHPLC-HRMS system to compare retention times (RT) and fragment masses.
4.4 LC Conditions
ResolveMass uses C18 or C8 stationary phases with optimized gradients of water/acetonitrile modified with formic acid or TFA. Elevated column temperatures (e.g., 65 circ to 80 circ ) are employed to enhance resolution of hydrophobic peptide fragments.
4.5 Data Analysis
Software-assisted peak matching evaluates RT similarity and mass-to-charge (m/z) ratios. Intensity ratios of corresponding fragments are compared to ensure consistent process control.
4.6 Key Observations / Acceptance Criteria
The fingerprint must show 1:1 peak correspondence. Any “new peak” in the test sample digest that is not in the RLD must be flagged for structural identification and risk assessment.
5. Impurity Profiling by HRMS
Impurity profiling ensures all peptide-related impurities are identified and controlled within regulatory limits.
Failure to properly characterize impurities is a major cause of regulatory delays, as highlighted in Peptide Sameness Study Deficiencies.
5.1 Objective
The objective of impurity profiling in a Tirzepatide Sameness Study is to identify, characterize, and quantify all peptide-related substances to ensure they do not exceed RLD levels or introduce new immunogenicity risks.
5.2 Types of Expected Impurities
Common synthetic impurities include amino acid deletions (e.g., -Des-Gly), insertions, oxidations (specifically at Trp-25), and deamidations. We also monitor for incomplete deprotection adducts like +Fmoc.
5.3 Experimental Approach
We utilize targeted LC-HRMS screening to mine for known impurity masses within the 0.05% – 0.10% range. This approach allows for the detection of co-eluting impurities that UV detectors alone cannot resolve.
5.4 LC-HRMS Conditions
Analysis is performed on the Thermo Q Exactive Plus or SCIEX TripleTOF platforms. These high-resolution systems provide the mass accuracy needed to distinguish between near-isobaric species like deamidated vs. native peptides.
5.5 Data Processing
Automated workflows in UNIFI or waters_connect compare the impurity levels against pre-defined regulatory thresholds. New specified impurities are structurally elucidated using MS/MS fragmentation data.
5.6 Key Observations / Acceptance Criteria
- Each peptide-related impurity 0.10% must be identified.
- Total new impurities must not exceed 0.5%.
- Impurity levels in the test product should be equal to or lower than the RLD.
6. Intact Mass Analysis
6.1 Objective
The objective is to confirm the global molecular identity of the Tirzepatide molecule by measuring its intact mass within extremely tight tolerances.
6.2 Method
High-resolution ESI-MS is used to generate multiple charge states (e.g., [M+3H]^{3+} to [M+5H]^{5+}). These are then mathematically deconvoluted to determine the neutral monoisotopic mass.
6.3 Observations
The measured mass must match the theoretical value for Tirzepatide (approx. 4813.53 Da). This step acts as a high-level screen to ensure the acylation and all 39 amino acids are present.
7. NMR Characterization (Orthogonal Confirmation)
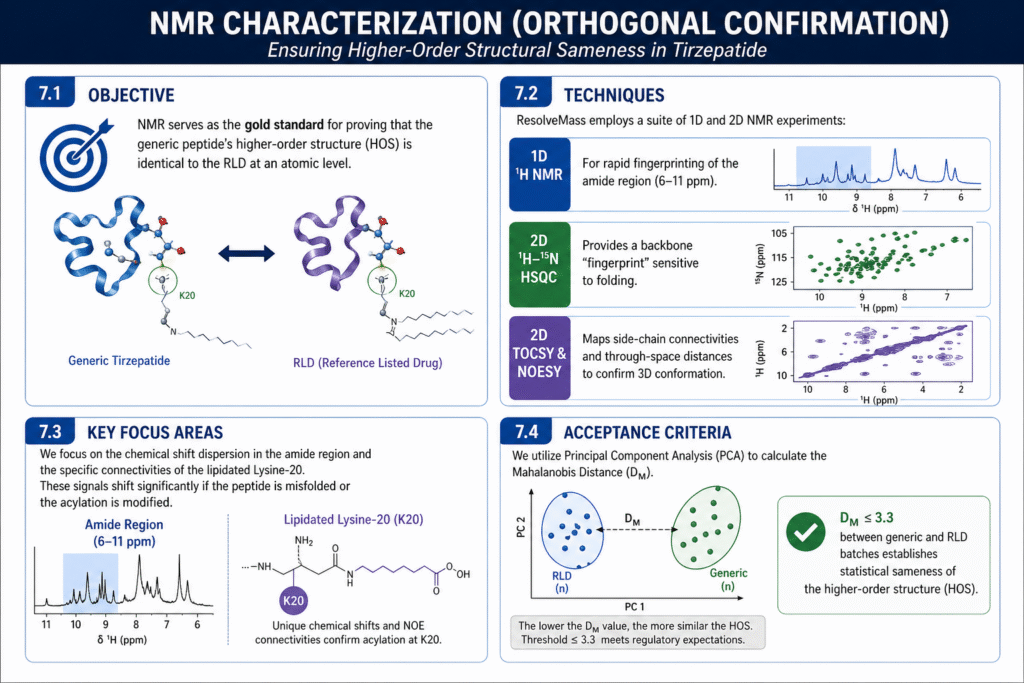
7.1 Objective
NMR serves as the gold standard for proving that the generic peptide’s higher-order structure (HOS) is identical to the RLD at an atomic level.
7.2 Techniques
ResolveMass employs a suite of 1D and 2D NMR experiments :
- 1D ^1H NMR: For rapid fingerprinting of the amide region (6-11 ppm).
- 2D ^1H-^{15}N HSQC: Provides a backbone “fingerprint” sensitive to folding.
- 2D TOCSY & NOESY: Maps side-chain connectivities and through-space distances to confirm 3D conformation.
7.3 Key Focus Areas
We focus on the chemical shift dispersion in the amide region and the specific connectivities of the lipidated Lysine-20. These signals shift significantly if the peptide is misfolded or the acylation is modified.
7.4 Acceptance Criteria
We utilize Principal Component Analysis (PCA) to calculate the Mahalanobis Distance (D_M). A D_M 3.3 between generic and RLD batches establishes statistical sameness of the HOS.
8. Comparative Sameness Assessment (RLD vs Test)
A successful Tirzepatide Sameness Study concludes with a comprehensive statistical comparison of at least three batches of the generic product vs. three batches of the RLD. This assessment includes PCA plots of NMR data, overlaid peptide maps, and side-by-side impurity tables. This holistic view proves that any minor manufacturing variations are within the range of the innovator’s own lot-to-lot variability.
9. Regulatory Considerations for ANDA
Regulatory success depends on demonstrating identity, impurity control, and structural equivalence.
Companies offering Peptide Sameness Study Services in United States and Peptide Sameness Study Services in Canada must align with global expectations.
Under the FDA’s May 2021 guidance, synthetic peptides referencing rDNA drugs must demonstrate “identity” through analytical sameness. A primary hurdle for Tirzepatide ANDAs is the justification of “new” impurities; if a synthetic process introduces a variant not found in the rDNA-origin RLD, ResolveMass provides the structural and toxicological data needed to prove that such differences do not impact safety or immunogenicity.
10. Conclusion
The Tirzepatide Sameness Study is a multi-disciplinary effort that translates complex analytical data into regulatory proof of equivalence. By integrating HRMS sequencing, multi-enzymatic mapping, and multidimensional NMR, ResolveMass Laboratories Inc. ensures that every critical quality attribute of Tirzepatide is meticulously verified. As an expert partner in generic peptide development, ResolveMass provides the scientific rigor and data integrity required to achieve ANDA success and accelerate the delivery of high-quality, affordable therapeutics to patients.
Frequently Asked Questions:
A Tirzepatide Sameness Study is a comprehensive analytical evaluation conducted to demonstrate that a generic version of tirzepatide is structurally and functionally equivalent to the reference listed drug (RLD). It includes techniques like peptide sequencing, mapping, HRMS/LCMS, and NMR to confirm sameness.
Peptide sequencing confirms the exact 39-amino acid structure of tirzepatide. It ensures there are no substitutions, deletions, or modifications compared to the RLD, which is critical for regulatory approval and therapeutic equivalence.
Peptide mapping generates a structural fingerprint by breaking the peptide into fragments and analyzing them using LC-MS. It helps detect minor variations, confirm sequence integrity, and ensure batch-to-batch consistency.
HRMS/LCMS identifies and quantifies low-level impurities such as truncated peptides, oxidized variants, and deamidated forms. It provides high-resolution, accurate mass data essential for meeting strict regulatory requirements.
NMR is used to confirm the higher-order structure (HOS) of tirzepatide at an atomic level. It ensures that the generic peptide folds and behaves structurally the same as the RLD, which is critical for biological activity.
Tirzepatide analysis is challenging due to its long peptide chain, presence of non-natural amino acids (Aib), and lipid (C20 fatty diacid) modification. These features require advanced and orthogonal analytical techniques for complete characterization.
Reference
- Belančić A, Antza C, Poutachidis A, Palaska S, Gkrinia EM, Faour AK, Sener YZ, Sener S, Sultana R. Comparative efficacy of GLP‐1 RA, tirzepatide and SGLT‐2 inhibitors in metabolic liver disease: A network meta‐analysis. British Journal of Clinical Pharmacology. 2026 Feb 17.https://bpspubs.onlinelibrary.wiley.com/doi/abs/10.1002/bcp.70492
- Crisafulli S, Alkabbani W, Paik JM, Bykov K, Tavakkoli A, Glynn RJ, Htoo PT, Yu EW, Trifirò G, Wexler DJ, Patorno E. Comparative gastrointestinal safety of dulaglutide, semaglutide, and tirzepatide in adults with type 2 diabetes. Annals of internal medicine. 2026 Jan;179(1):1-1.https://www.acpjournals.org/doi/abs/10.7326/ANNALS-25-01724
- Cariou B, Linge J, Neeland IJ, Dahlqvist Leinhard O, Petersson M, Fernández Landó L, Bray R, Rodríguez Á. Effect of tirzepatide on body fat distribution pattern in people with type 2 diabetes. Diabetes, Obesity and Metabolism. 2024 Jun;26(6):2446-55.https://dom-pubs.onlinelibrary.wiley.com/doi/abs/10.1111/dom.15566
- Henney AE, Riley DR, Anson M, Heague M, Hernadez G, Alam U, Craig S, Cuthbertson DJ. Comparative efficacy of tirzepatide, liraglutide, and semaglutide in reduction of risk of major adverse cardiovascular events in patients with obstructive sleep apnea and type 2 diabetes: real-world evidence. Annals of the American Thoracic Society. 2025 Jul;22(7):1042-52.https://academic.oup.com/annalsats/article-abstract/22/7/1042/8381937


