Introduction:
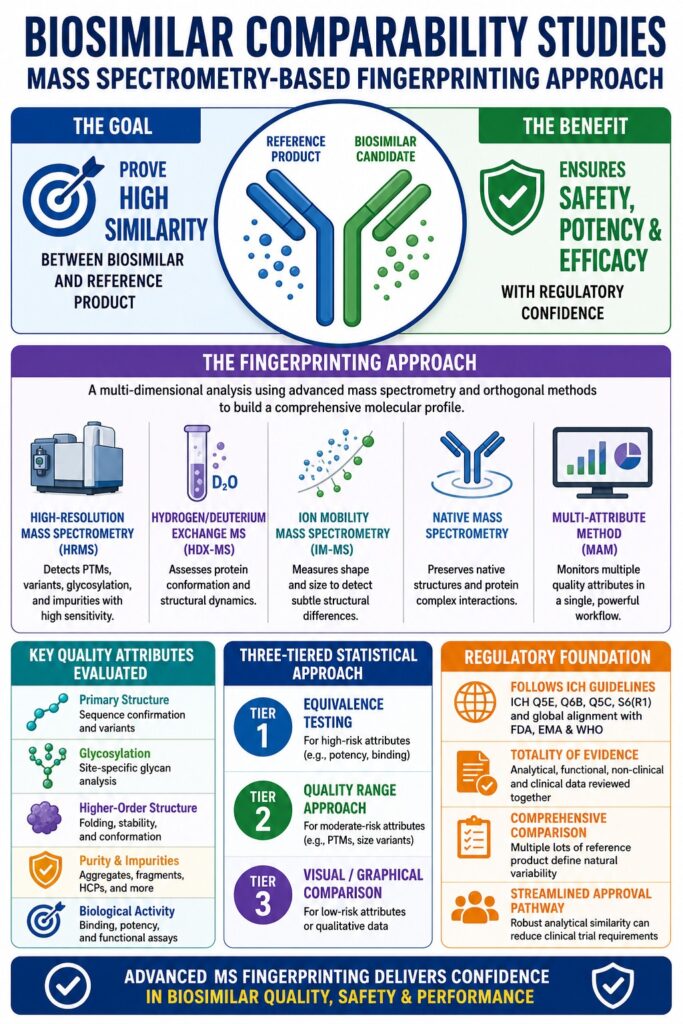
Biosimilar Comparability Studies use an advanced fingerprinting strategy to prove that a follow-on biologic is highly similar to its reference product. Instead of relying on one analytical test, this strategy creates a detailed and multidimensional molecular profile of the biosimilar. The process combines high-resolution mass spectrometry (HRMS), hydrogen-deuterium exchange mass spectrometry (HDX-MS), and multi-attribute methods (MAM) to evaluate structural and functional similarity at an extremely high level of precision. These technologies help detect even the smallest variations in post-translational modifications (PTMs), glycosylation patterns, or higher-order structures that could potentially affect therapeutic performance.
Modern biosimilar development depends heavily on analytical characterization because biologic drugs are naturally complex and heterogeneous. Even slight changes in manufacturing conditions can alter the molecular structure of a biologic product. Therefore, fingerprinting studies are designed to confirm that any observed differences remain within an acceptable range and do not impact clinical safety, potency, purity, or efficacy. This analytical approach has significantly reduced dependence on extensive clinical trials by establishing confidence through robust structural and functional evidence.
The use of mass spectrometry-based fingerprinting has transformed the biosimilar industry by enabling deeper molecular insight than traditional analytical methods. Regulatory agencies now prioritize analytical similarity as the foundation of biosimilar approval pathways. As a result, pharmaceutical developers increasingly invest in sophisticated characterization workflows that improve comparability assessments and accelerate product development timelines.
Explore our specialized services for fingerprinting and analytical similarity: Analytical Services for Generic Drug Development
Share via:
The Regulatory Framework of Biosimilar Comparability Studies
Biosimilar Comparability Studies are designed to establish analytical similarity between a proposed biosimilar and its originator biologic through comprehensive evaluation of critical quality attributes (CQAs). These studies follow the “totality of evidence” framework, where analytical, functional, non-clinical, and clinical data are collectively reviewed to determine biosimilarity. Among these components, analytical characterization remains the most important and sensitive element because it directly measures molecular similarity between products.
Under ICH Q5E guidance, comparability assessments are intended to demonstrate that manufacturing changes or biosimilar development processes do not negatively affect predefined attributes related to safety, purity, and potency. Regulatory agencies require manufacturers to prove that the biosimilar maintains the same structural and functional characteristics as the reference product. This includes detailed evaluation of protein sequence, glycosylation, aggregation, charge heterogeneity, and biological activity.
The international regulatory landscape is harmonized across agencies such as the FDA, EMA, and WHO. These organizations apply the principles of ICH Q6B to define the physicochemical and biological specifications necessary for biotechnology-derived products. Biosimilar manufacturers must perform extensive analytical characterization before regulatory authorities consider reducing clinical trial requirements. A highly comprehensive analytical package demonstrating near-identical structural and functional behavior can streamline development programs and accelerate market approval.
Regulatory agencies now strongly encourage “fingerprint-like” analysis, where numerous product attributes are assessed simultaneously using highly sensitive orthogonal methods. Sponsors typically analyze multiple batches of the reference product, often around 10 or more lots, to establish an acceptable statistical variability range. This approach ensures that natural variability of the originator product is fully understood before comparing the biosimilar candidate.
Ensure your project meets global standards with expert guidance: Regulatory Support for Generic Drug Development
ICH Guidelines and Global Regulatory Alignment
The foundation of biosimilar comparability assessment is ICH Q5E, which provides harmonized guidance for evaluating the impact of manufacturing changes on biological products. Although originally intended for in-house manufacturing modifications, its scientific principles are widely applied to biosimilarity assessment between products manufactured by different companies. The guideline establishes the scientific logic needed to demonstrate that no clinically meaningful differences exist between the biosimilar and the reference product.
| ICH Guideline | Focus Area | Application in Biosimilarity |
| ICH Q5E | Comparability of biotechnological/biological products subject to changes. | Establishes the logic for proving “no clinically meaningful difference” after a change. |
| ICH Q6B | Specifications for biotechnological and biological products. | Defines test procedures for physicochemical properties, purity, and activity. |
| ICH Q5C | Stability testing of biotechnological/biological products. | Ensures the biosimilar maintains its quality profile throughout its shelf life. |
| ICH S6 (R1) | Preclinical safety evaluation. | Guides the transition from analytical to non-clinical and clinical assessments. |
The Three-Tiered Statistical Approach
Analytical similarity evaluation is performed using a structured tiered statistical approach rather than a simple pass/fail comparison. This framework categorizes quality attributes based on their clinical importance and associated risk. Each tier applies a different statistical method depending on the potential impact of the attribute on product safety and efficacy.
| Tier | Statistical Method | Application |
| Tier 1 | Equivalence Testing | Reserved for highest-risk attributes like potency and target binding. |
| Tier 2 | Quality Range Approach | Used for moderate-risk attributes like PTMs and size variants. |
| Tier 3 | Visual/Graphical Comparison | Applied to lowest-risk attributes or qualitative raw data. |
The Fingerprinting Paradigm: Advanced Bioanalytical Characterization
The fingerprinting paradigm represents a major advancement in Biosimilar Comparability Studies because it evaluates the biologic as an integrated molecular system rather than focusing on isolated tests. By combining multiple orthogonal analytical technologies, scientists can create a comprehensive molecular profile that captures structural, physicochemical, and functional characteristics simultaneously.
Orthogonal analytical methods use different scientific principles to examine the same attribute, improving confidence in the final comparability conclusion. For example, higher-order structure can be assessed through spectroscopic methods such as circular dichroism (CD) and Fourier-transform infrared spectroscopy (FTIR), while mass spectrometry-based methods like HDX-MS and ion mobility mass spectrometry (IM-MS) provide deeper insight into protein folding and conformational dynamics.
Bioanalytical characterization begins with confirmation of the complete amino acid sequence and expands into evaluation of PTMs such as oxidation, deamidation, glycation, and glycosylation. Since biologics are inherently heterogeneous, the objective is not to achieve absolute uniformity but to demonstrate that the biosimilar falls within the established variability range of the reference product.
The fingerprinting approach also improves understanding of structure-function relationships. Subtle molecular changes can influence immunogenicity, stability, pharmacokinetics, or biological activity. Therefore, comprehensive characterization is essential to ensure that the biosimilar behaves consistently under clinical conditions.
Learn how mass spectrometry is used to define biosimilar profiles: Biosimilar Characterization Using Mass Spectrometry
Selection of Critical Quality Attributes (CQAs)
The selection of Critical Quality Attributes (CQAs) depends on the mechanism of action (MOA) and clinical behavior of the biologic molecule. CQAs are properties that directly influence product quality, safety, efficacy, or immunogenicity. During biosimilar development, manufacturers must identify and prioritize these attributes for detailed analytical comparison.
For monoclonal antibodies (mAbs), glycosylation is often classified as a Tier 1 or Tier 2 attribute because it influences antibody-dependent cellular cytotoxicity (ADCC), complement activation, serum half-life, and structural stability. Even small changes in glycan composition may alter therapeutic activity or immunogenicity, making accurate glycan analysis critical.
Primary structure characterization includes amino acid sequence confirmation and N-terminal/C-terminal sequencing. Physicochemical characterization evaluates molecular weight, isoform distribution, and electrophoretic behavior. Higher-order structure analysis examines secondary, tertiary, and quaternary protein folding patterns to confirm conformational integrity.
Purity assessments focus on detecting aggregates, fragments, process-related impurities, and host cell proteins (HCPs). Biological activity studies evaluate target binding, receptor activation, and potency relevant to the therapeutic mechanism of action. Together, these CQAs provide a comprehensive framework for biosimilarity evaluation.
Analyze complex glycan profiles to ensure therapeutic efficacy: Glycosylation Analysis of Biosimilars
High-Resolution Mass Spectrometry (HRMS) Platforms
High-resolution mass spectrometry is the core technology behind modern Biosimilar Comparability Studies because it provides exceptional mass accuracy, sensitivity, and resolving power. HRMS platforms can identify and quantify complex proteoforms, PTMs, and low-abundance variants that may not be detectable using conventional analytical techniques.
The biopharmaceutical industry has gradually transitioned from traditional triple quadrupole and time-of-flight (TOF) systems toward Orbitrap-based instruments. Orbitrap platforms offer superior mass resolution, improved stability, and enhanced native-MS capabilities, making them highly suitable for detailed biologic characterization.
Recent advancements in Orbitrap technology allow accurate mass analysis beyond 8,000 m/z, enabling characterization of intact monoclonal antibodies under native conditions. These systems can resolve individual glycoforms with high precision, providing detailed molecular fingerprints that support biosimilarity assessment.
Fourier Transform-based signal detection in Orbitrap systems improves signal clarity by reducing matrix interference and background noise. This enhanced sensitivity is especially important for identifying low-level impurities, degradation products, or rare structural variants that may affect product quality or safety.
Orbitrap vs. Q-TOF in Biotherapeutic Analysis
Orbitrap and Q-TOF instruments are both widely used in biotherapeutic analysis, but they differ in several important technical capabilities. Historically, Q-TOF systems were preferred for large biomolecule analysis due to their broad mass range and fast acquisition speed. However, modern Orbitrap systems now provide comparable or superior performance for many biosimilar applications.
| Feature | Orbitrap (e.g., Q Exactive / Orbitrap Ascend) | Q-TOF (e.g., Agilent 6550 / Xevo G3) |
|---|---|---|
| Mass Accuracy | <1–3 ppm | <1–5 ppm |
| Resolution | Up to 280,000 (at m/z 200) | Typically 40,000–60,000 |
| Native MS | Exceptional; BioPharma Option allows native-like acquisition. | Good for average mass analysis. |
| Quantitative Ability | Comparable to triple quadrupoles with HRAM benefits. | Strong for rapid peptide mapping. |
| Robustness | High stability; system often calibrated once a week. | May require more frequent expert tuning for high-mass work. |
In biosimilar characterization, the choice between Orbitrap and Q-TOF often depends on the analytical objective. Many laboratories use both platforms in complementary workflows to maximize coverage, sensitivity, and confidence in comparability assessments.
Instrumental Parameters for HRAM Analysis
Successful high-resolution accurate mass (HRAM) analysis depends on careful optimization of instrumental parameters. For intact protein analysis, settings must preserve large molecular ions while maintaining sufficient sensitivity and resolution for accurate mass determination.
Orbitrap-based workflows often use resolution settings ranging from 15,000 to 240,000 depending on analytical requirements. Automatic Gain Control (AGC) values may reach up to 5 × 10^6 to ensure adequate ion populations for deconvolution and quantitative analysis. These optimized conditions improve data reproducibility and spectral quality.
Source parameters such as sheath gas flow, auxiliary gas flow, spray voltage, and ion transfer conditions are standardized to achieve reproducible ionization using Heated Electrospray Ionization (HESI). Consistent source conditions are particularly important when comparing biosimilar batches across multiple analytical runs or laboratories.
Standardization of HRAM parameters also supports inter-laboratory reproducibility and regulatory compliance. Consistent analytical procedures ensure reliable comparison of structural data throughout product development and lifecycle management.
Analytical Strategies: Bottom-Up, Middle-Down, and Top-Down
Mass spectrometry-based fingerprinting uses three major analytical strategies: Bottom-Up, Middle-Down, and Top-Down proteomics. Each strategy provides different levels of structural detail and molecular context.
Bottom-Up: Detailed Peptide Mapping
Bottom-Up proteomics is considered the workhorse of biosimilar characterization. In this method, proteins are enzymatically digested into smaller peptides using proteases such as trypsin before LC-MS/MS analysis. This approach provides extremely high sequence coverage and enables site-specific identification of PTMs.
Bottom-Up workflows are highly effective for detecting deamidation, oxidation, glycation, and sequence variants. Because peptides are analyzed individually, researchers can identify the exact amino acid residue associated with each modification. This level of detail is essential for demonstrating structural similarity between biosimilars and reference products.
Despite its advantages, Bottom-Up analysis requires extensive sample preparation, which can introduce artifacts such as artificial oxidation or deamidation during digestion. Careful method optimization and control experiments are necessary to minimize analytical variability.
Identify site-specific modifications with precision peptide mapping: Peptide Mapping in Biosimilars
Middle-Down: Subunit Characterization
Middle-Down and Middle-Up approaches divide large proteins into intermediate-sized subunits using enzymes like IdeS followed by reduction of disulfide bonds. This strategy simplifies analysis while preserving more structural context than Bottom-Up workflows.
Subunit analysis is particularly useful for monoclonal antibodies because it enables characterization of light chain (LC), Fd’, and Fc/2 fragments individually. Researchers can rapidly evaluate glycosylation patterns, oxidation, and structural heterogeneity at the subunit level.
Middle-Down workflows are often faster than peptide mapping and require less extensive sample preparation. They are also beneficial for dilute or unstable samples where complete digestion may be difficult to achieve consistently.
Top-Down: Intact Mass Analysis
Top-Down proteomics analyzes intact proteins without enzymatic digestion, preserving the complete proteoform structure. This approach provides a global overview of PTM combinations and glycoform distributions on individual molecules.
One major advantage of Top-Down analysis is the reduced risk of preparation-induced artifacts because proteins remain close to their native state. The technique is particularly useful for detecting major structural shifts or glycan profile changes that may impact biosimilarity.
However, Top-Down workflows can be analytically challenging due to the complexity of intact protein fragmentation and data interpretation. Advanced fragmentation methods such as ultraviolet photodissociation (UVPD) and electron-transfer dissociation (ETD) are often required for detailed structural analysis.
Achieve a global overview of your protein structure: Intact Mass Analysis of Biosimilars
Higher-Order Structure (HOS) Characterization via MS
Higher-order structure (HOS) characterization is mandatory in Biosimilar Comparability Studies because protein folding directly influences biological activity, stability, and immunogenicity. Even when primary sequences are identical, differences in tertiary or quaternary structure may alter therapeutic performance.
Mass spectrometry-based methods have become highly valuable for HOS analysis because they provide dynamic and structural information beyond conventional spectroscopy. These approaches help evaluate protein conformational stability, folding behavior, and molecular interactions under native-like conditions.
HOS characterization is particularly important for monoclonal antibodies, fusion proteins, and other complex biologics where conformational integrity determines receptor binding and effector function. Advanced MS techniques can identify subtle conformational changes that may not be visible through standard physicochemical assays.
Combining multiple orthogonal HOS techniques improves analytical confidence and strengthens regulatory submissions. This integrated approach provides a more complete understanding of biosimilar structural comparability.
Hydrogen/Deuterium Exchange Mass Spectrometry (HDX-MS)
HDX-MS is considered the gold standard for studying protein structural dynamics and conformational behavior. The technique measures the exchange rate of backbone amide hydrogens with deuterium in a D₂O-containing buffer. Exchange rates depend on hydrogen bonding and solvent accessibility, making HDX-MS highly sensitive to protein folding and flexibility.
In Biosimilar Comparability Studies, HDX-MS helps confirm that the biosimilar exhibits conformational dynamics similar to the reference product. Matching exchange profiles indicate comparable structural stability and folding behavior. This information is critical for confirming biological function and minimizing immunogenicity risk.
HDX-MS also supports epitope mapping by identifying interaction regions between therapeutic proteins and their binding targets. Additionally, the technique can detect early stages of protein unfolding or aggregation before visible degradation occurs.
Researchers at NIST have improved HDX-MS reproducibility through subzero chromatography workflows performed at approximately -30°C. These optimized methods reduce back-exchange and improve precision across laboratories, supporting more reliable biosimilar characterization.
Ion Mobility-Mass Spectrometry (IM-MS)
Ion Mobility-Mass Spectrometry adds another analytical dimension by separating ions based on shape, size, and charge while they travel through a gas-filled chamber. The technique generates Collision Cross Section (CCS) measurements that reflect the three-dimensional structure of ions in the gas phase.
Small changes in CCS values may indicate conformational differences caused by sequence variation, stress conditions, pH changes, or altered glycosylation. IM-MS therefore provides valuable information about higher-order structure and molecular flexibility.
The Mason-Schamp equation describes ion mobility:
K=3ze16N(2πμkBT)1/21ΩK = \frac{3ze}{16N} \left( \frac{2\pi}{\mu k_B T} \right)^{1/2} \frac{1}{\Omega}K=16N3ze(μkBT2π)1/2Ω1
In this equation, Ω represents the Collision Cross Section (CCS). Because CCS is directly related to ion conformation, IM-MS can identify subtle structural changes that may not be detectable through intact mass measurements alone.
IM-MS is increasingly integrated into biosimilar workflows because it complements HRMS and HDX-MS data. Together, these techniques provide comprehensive structural characterization at multiple levels of molecular organization.
Native Mass Spectrometry
Native Mass Spectrometry preserves non-covalent interactions and native-like protein structures during gas-phase analysis. This is achieved using volatile buffers and physiological pH conditions compatible with electrospray ionization.
Native MS enables direct observation of tertiary and quaternary protein structures, protein-protein interactions, and ligand-binding stoichiometry. The method is highly valuable for studying intact monoclonal antibodies, antibody-drug conjugates, and protein complexes.
A key parameter in native MS is the Charge State Distribution (CSD). Properly folded proteins typically exhibit narrow charge state distributions shifted toward higher m/z regions. Changes in CSD patterns may indicate partial unfolding, aggregation, or loss of structural integrity.
Because native MS minimizes sample perturbation, it provides a more biologically relevant view of protein structure and behavior. This makes it an increasingly important tool for biosimilar comparability assessment and structural confirmation.
The Multi-Attribute Method (MAM) in GxP Environments
The Multi-Attribute Method (MAM) is an LC-MS-based peptide mapping strategy developed to replace multiple traditional analytical assays with a single, highly informative workflow. MAM simultaneously monitors numerous product quality attributes (PQAs), improving efficiency and analytical consistency throughout biologic development and quality control.
| Feature | Conventional Assays | Multi-Attribute Method (MAM) |
| PQA Coverage | Single attribute per test (e.g., glycans only). | Multiple attributes (glycans, deamidation, oxidation). |
| Site Specificity | Low; often provides general averages. | High; residue-specific quantification. |
| Unknown Detection | Limited to specific method range. | Comprehensive New Peak Detection (NPD). |
| Throughput | Multiple assays required; time-consuming. | Single injection; streamlined workflow. |
The adoption of MAM in GxP environments has increased significantly due to advances in compliance-ready software such as Byosphere and UNIFI. These systems support automated reporting, electronic records, and 21 CFR Part 11 compliance, improving data integrity and operational efficiency.
Validate similarity with advanced LC-MS workflows: Prove Biosimilarity Using LC-MS
Glycosylation and PTM Site-Specificity
Glycosylation is one of the most critical quality attributes in Biosimilar Comparability Studies because it directly influences pharmacokinetics, immunogenicity, serum stability, and biological activity. Regulatory agencies require detailed site-specific glycosylation analysis rather than generalized glycan averages.
Traditional released glycan analysis removes glycans from the protein backbone, eliminating information about glycosylation attachment sites. In contrast, glycopeptide-based MS analysis preserves the peptide-glycan connection and allows precise site-specific characterization.
Fc-region glycans are particularly important because they influence antibody-dependent cellular cytotoxicity (ADCC) and complement activation. Fab-region glycans can affect antigen binding and molecular stability. Accurate characterization of both regions is essential for establishing biosimilarity.
Advanced fragmentation techniques such as ETD and UVPD preserve labile PTMs during peptide fragmentation, enabling precise localization of glycosylation sites. These technologies improve confidence in glycoform identification and support detailed structure-function analysis.
Map and quantify post-translational modifications accurately: Post-Translational Modifications (PTMs) in Biosimilars
Software and Informatics for Biosimilarity
The enormous volume of data generated during Biosimilar Comparability Studies requires sophisticated software and informatics solutions. Modern analytical platforms integrate automated workflows for deconvolution, peptide mapping, glycoform analysis, and statistical comparison.
Software tools such as Byos, BioPharma Finder, and waters_connect provide comprehensive ecosystems for biotherapeutic characterization. These systems automate sequence coverage calculations, PTM quantification, and comparative fingerprint analysis across multiple batches.
Dashboard visualization tools allow researchers to monitor trends and rapidly identify attributes that fall outside predefined variability ranges. Automated statistical evaluation improves consistency and reduces manual interpretation errors.
Data integrity and regulatory compliance are also critical considerations. Biosimilar analytical platforms must comply with 21 CFR Part 11 requirements, including electronic audit trails, secure data management, and validated reporting systems. These informatics solutions play a central role in supporting global regulatory submissions.
Identify and profile potential impurities in your biosimilar: Impurity Profiling of Biosimilars
Conclusion
Biosimilar Comparability Studies have evolved into highly sophisticated, mass spectrometry-driven fingerprinting programs that provide unprecedented insight into complex biotherapeutic molecules. Advanced analytical technologies such as Orbitrap HRMS, HDX-MS, IM-MS, native MS, and MAM collectively enable comprehensive structural and functional characterization with remarkable sensitivity and precision.
The integration of orthogonal analytical techniques allows developers to demonstrate the “essential sameness” of biosimilars while ensuring patient safety, therapeutic efficacy, and regulatory confidence. This analytical rigor forms the scientific foundation of the “totality of evidence” approach used by global health authorities.
As biosimilar development continues to expand worldwide, mass spectrometry-based fingerprinting will remain central to regulatory approval strategies. Continuous improvements in instrumentation, software, and data analysis are expected to further enhance analytical sensitivity, streamline development workflows, and reduce time-to-market for new biosimilar therapies.
Ultimately, these advanced comparability approaches support broader patient access to affordable biologic therapies while maintaining the highest standards of quality, safety, and clinical performance.
Get started on your biosimilar journey with our specialized development services: Formulation Development for Generic Drug Development
For specialized support in navigating these complex analytical requirements or to leverage state-of-the-art MS fingerprinting for your biosimilar program, please connect with our technical team.
Contact Us: Resolve Mass Contact Page
Frequently Asked Questions
Regulatory agencies prefer fingerprint-like analysis because it evaluates a broad spectrum of molecular and functional attributes simultaneously. This approach offers a highly sensitive and comprehensive comparison between the biosimilar and the reference product. By examining multiple quality parameters together, regulators gain stronger confidence that there are no clinically meaningful differences. It also supports reduced reliance on extensive clinical trials by providing detailed analytical evidence of similarity.
Mass spectrometry identifies site-specific glycosylation through glycopeptide analysis, where glycans remain attached to their original amino acid residues during testing. This enables precise determination of both the glycan structure and its exact attachment location on the protein. Such detailed mapping is important because glycosylation at different regions can affect therapeutic activity differently. The technique also improves understanding of protein stability, immunogenicity, and biological function.
ICH Q5E provides the scientific framework used to assess whether manufacturing changes impact the quality, safety, or efficacy of a biological product. In biosimilar development, these principles are applied to compare products made by different manufacturers. The guideline helps establish comparability by focusing on analytical consistency and critical quality attributes. It also supports a risk-based approach for regulatory evaluation and lifecycle management.
Orbitrap mass spectrometers offer extremely high mass resolution and excellent mass accuracy, making them highly effective for biosimilar characterization. These systems can resolve complex glycoforms and detect subtle molecular differences that may not be visible with conventional instruments. Orbitrap platforms also provide superior native mass spectrometry performance for analyzing intact proteins and protein complexes. Their high stability and sensitivity improve data reproducibility across analytical workflows.
HDX-MS evaluates higher-order structure (HOS) by measuring hydrogen-to-deuterium exchange rates within the protein backbone. These exchange patterns reflect protein folding, conformational flexibility, and solvent accessibility. When the biosimilar and reference product show matching exchange profiles, it indicates comparable structural dynamics and folding behavior. This technique is especially useful for confirming conformational stability and minimizing immunogenicity concerns.
The three-tier statistical approach categorizes quality attributes based on their clinical importance and associated risk. Tier 1 uses equivalence testing for high-risk attributes such as potency and receptor binding. Tier 2 applies quality range analysis for moderate-risk attributes like glycosylation and charge variants. Tier 3 relies on visual or graphical comparison for lower-risk qualitative data and supportive analytical observations.
Collision Cross Section (CCS) is an important parameter in Ion Mobility-Mass Spectrometry because it reflects the three-dimensional shape and size of an ion. Even if two proteins have identical masses, differences in CCS values can reveal structural or conformational variations. CCS measurements therefore help confirm higher-order structural equivalence between biosimilars and reference products. This information is valuable for detecting subtle folding changes that may influence therapeutic performance.
Reference:
- Berkowitz, S. A., Engen, J. R., Mazzeo, J. R., & Jones, G. B. (2012). Analytical tools for characterizing biopharmaceuticals and the implications for biosimilars. Nature Reviews Drug Discovery, 11(7), 527–540. https://doi.org/10.1038/nrd3746
- U.S. Food and Drug Administration. (2020). Statistical approaches to evaluate analytical similarity: Guidance for industry. U.S. Department of Health and Human Services. https://www.fda.gov/media/135612/download
- Hines, K. M., Ross, D. H., Davidson, K. L., Bush, M. F., & Xu, L. (2017). Large-scale structural characterization of drug and drug-like compounds by high-throughput ion mobility-mass spectrometry. Analytical Chemistry, 89(17), 9023–9030. https://doi.org/10.1021/acs.analchem.7b01709
- Chow, S.-C., Song, F., & Bai, H. (2016). Analytical similarity assessment in biosimilar studies. The AAPS Journal, 18(3), 670–677. https://doi.org/10.1208/s12248-016-9882-5
- European Medicines Agency. (2014). Guideline on similar biological medicinal products (Rev. 1) (CHMP/437/04 Rev. 1). European Medicines Agency. https://www.ema.europa.eu/en/similar-biological-medicinal-products-scientific-guideline
- U.S. Food and Drug Administration. (2017). Considerations in demonstrating interchangeability with a reference product: Guidance for industry. U.S. Department of Health and Human Services. https://www.fda.gov/media/105013/download



