Introduction: The Analytical Importance of Impurity Profiling of Biosimilars
Impurity Profiling of Biosimilars is one of the most important steps in modern biosimilar development. Regulatory agencies such as the FDA and EMA require manufacturers to prove that a biosimilar closely matches the reference biologic not only in structure but also in impurity profile and sequence consistency. Even very small sequence variants, sometimes as low as 0.1%, can raise regulatory concerns because they may affect safety, efficacy, or immunogenicity.
Unlike traditional small-molecule drugs, biosimilars are produced using living cell systems. Because of this, small variations can occur during cell culture, protein expression, purification, or storage. Factors such as cell line selection, media composition, bioreactor settings, and purification conditions can all influence the final product quality. For this reason, detailed impurity characterization is essential for biosimilar approval and lifecycle management.
High-resolution mass spectrometry (HRMS) has become a leading analytical tool for Impurity Profiling of Biosimilars because it provides highly accurate mass measurements with exceptional sensitivity. HRMS helps scientists identify low-level host cell proteins (HCPs), sequence variants, glycoforms, and unexpected modifications with high confidence. The technology also supports both qualitative and quantitative analysis within a single workflow, improving efficiency and regulatory reliability.
Explore our specialized services: Biosimilar Characterization Using Mass Spectrometry
At ResolveMass Laboratories Inc., specialized HRMS workflows are developed and validated specifically for biosimilar impurity and sequence variant profiling. These workflows are aligned with global regulatory expectations for IND, BLA, and MAA submissions. By combining advanced instrumentation, regulatory expertise, and modern bioinformatics, ResolveMass Laboratories supports biosimilar developers with high-quality analytical comparability data.
Learn how to Prove Biosimilarity Using LC-MS
Share via:
📋 Article Summary
- High-resolution mass spectrometry (HRMS) plays a central role in the detection, characterization, and quantification of impurities and sequence variants in biosimilar products.
- Effective Impurity Profiling of Biosimilars relies on multiple complementary MS techniques, including peptide mapping, intact mass analysis, and native mass spectrometry, to confirm similarity with the reference biologic.
- Sequence variants caused by translational errors, genetic instability, or cell culture stress are closely monitored by regulatory authorities and must remain within accepted FDA, EMA, and ICH limits.
- Advanced technologies such as Orbitrap HRMS, ion mobility spectrometry, and data-independent acquisition (DIA) have significantly improved analytical sensitivity, accuracy, and workflow efficiency.
- Global regulatory frameworks, including ICH Q6B and biosimilar guidance from the FDA and EMA, require extensive evaluation of both process-derived and product-related impurities.
- ResolveMass Laboratories Inc. provides specialized HRMS-based analytical workflows tailored for biosimilar characterization, comparability assessments, and regulatory submission support.
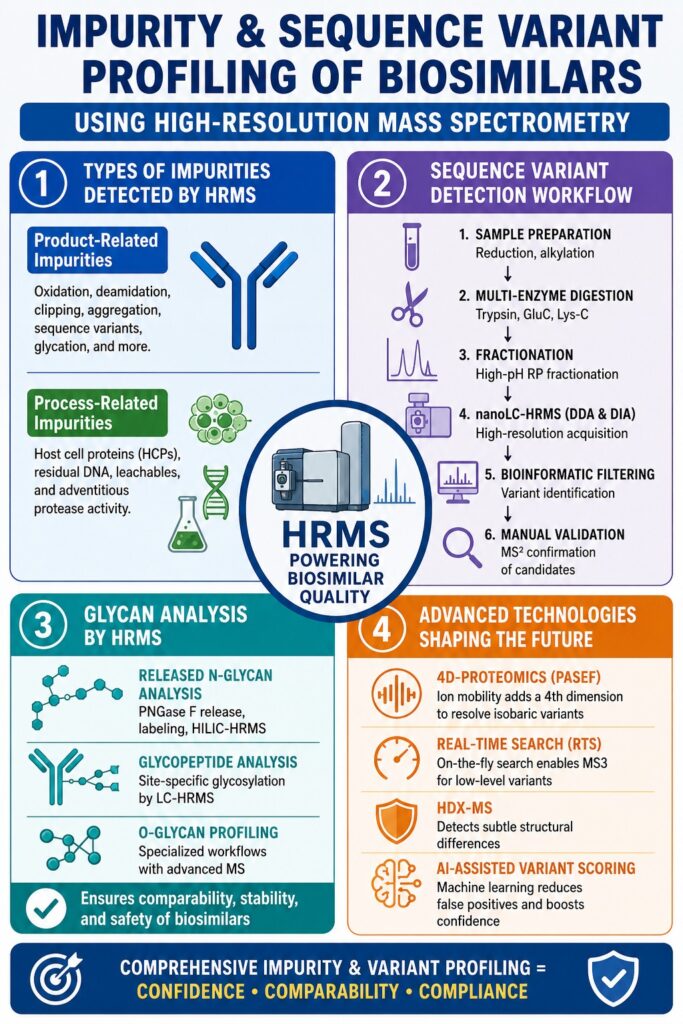
1. Classifying Impurities in Biosimilars: A Mass Spectrometry Perspective
Biosimilar impurities are generally grouped into two categories:
- Product-related impurities
- Process-related impurities
Product-related impurities originate from the biologic molecule itself, while process-related impurities come from manufacturing materials, host cells, or production environments. HRMS enables precise molecular-level identification and quantification of both impurity types.
International guidelines such as ICH Q6B require both qualitative and quantitative impurity assessment. Manufacturers must establish impurity thresholds, evaluate possible clinical impact, and demonstrate consistency across production lots. As a result, Impurity Profiling of Biosimilars has become a critical quality requirement throughout development and commercialization.
See our full range of Analytical Services for Generic Drug Development
Product-Related Impurities Detectable by HRMS
| Impurity Type | MS Detection Strategy | Typical Detection Limit |
|---|---|---|
| Oxidized methionine/tryptophan variants | Peptide mapping + MS/MS | 0.05–0.1% |
| Deamidated asparagine variants | High-pH peptide mapping + HRMS | 0.1% |
| Clipped/truncated species | Intact mass + SEC-MS | 0.5% |
| Aggregates (non-covalent) | Native MS / nSEC-MS | 0.1% |
| Sequence variants (amino acid substitutions) | DIA peptide mapping | 0.05% |
| Glycation adducts | Glycopeptide enrichment + HRMS | 0.1% |
These impurities may affect the stability, potency, and safety of biosimilars. Oxidation and deamidation can reduce protein stability, while aggregation may increase immunogenicity risk. Accurate detection of low-level modifications is therefore essential.
Process-Related Impurities Detectable by HRMS
Host Cell Proteins (HCPs)
LC-MS/MS with DIA workflows is commonly used to identify and quantify host cell proteins using large spectral libraries. HRMS can detect contaminants at sub-ppm levels with greater specificity than traditional ELISA methods.
Residual DNA
Residual DNA is typically measured using qPCR methods, but LC-MS can provide additional confirmation through nucleobase analysis in concentrated samples.
Leachables and Extractables
Mass spectrometry helps identify contaminants from tubing, plasticizers, filters, or silicone oils used during manufacturing. These substances can impact product quality and patient safety.
Adventitious Protease Activity
Unexpected protease activity may create characteristic clipping patterns that are visible during intact mass analysis. Monitoring these degradation products helps protect product integrity and shelf-life stability.
2. Sequence Variant Profiling in Impurity Profiling of Biosimilars
Sequence variants are amino acid substitutions that occur because of translational errors or cellular stress during protein production. Regulatory agencies generally expect these variants to remain below approximately 0.1% because even low-level changes may affect biologic activity or immunogenicity.
One major analytical challenge is that some variants differ from native peptides by only 1 Da. Detecting these differences requires high-resolution instrumentation, optimized chromatography, and advanced bioinformatics.
Why Sequence Variants Matter
The FDA and EMA classify sequence heterogeneity as a critical quality attribute (CQA). A single amino acid substitution in a functional region of a monoclonal antibody can alter:
- Receptor binding
- Potency
- Pharmacokinetics
- Immunogenicity
- Thermal stability
Because biosimilars must closely match the reference product, manufacturers are required to thoroughly evaluate all detectable variants.
View our expertise in Peptide Mapping in Biosimilars
HRMS Workflow for Sequence Variant Detection
- Protein Reduction and Alkylation: The biologic is fully denatured using reagents such as DTT/IAA or TCEP to ensure equal accessibility of peptide bonds. This step improves digestion efficiency and minimizes structural bias during peptide mapping experiments.
- Multi-Enzyme Digestion: Parallel digestion using Trypsin, GluC, and Lys-C maximizes sequence coverage beyond 99%. Multiple digestion strategies are essential because missed cleavages or poorly ionizing peptides can conceal low-abundance variants.
- High-pH Reversed-Phase Fractionation: Peptides are separated at elevated pH to reduce ion suppression and improve detection sensitivity for trace-level variants. Fractionation also enhances chromatographic resolution and increases peptide identification rates during LC-MS analysis.
- nanoLC-HRMS Acquisition (DDA and DIA): Both data-dependent acquisition (DDA) and data-independent acquisition (DIA) methods are employed. DDA supports discovery-based identification, while DIA enables reproducible quantification across complex peptide populations. Orbitrap Astral and Q Exactive HF-X systems provide mass accuracy below 2 ppm for highly confident analysis.
- Bioinformatic Filtering: Advanced software platforms such as Byonic, Mascot, and Proteome Discoverer evaluate all possible amino acid substitutions against the reference sequence. Customized scoring thresholds and false-discovery controls are implemented to minimize false-positive identifications.
- Manual Validation of Candidates: All sequence variant candidates above approximately 0.03% abundance undergo manual MS2 spectral review for b/y-ion confirmation before final reporting. This additional verification step ensures analytical reliability and regulatory defensibility.
Quantification Approaches
- Label-Free Quantification (LFQ): LFQ compares peptide signal intensities between variant and wild-type peptides. While highly sensitive, quantitative accuracy may be affected by differences in peptide ionization efficiency.
- Stable Isotope Dilution (SID): This method introduces isotopically labeled reference peptides as internal standards, providing highly accurate quantitative measurements. SID approaches are frequently used in regulatory submissions requiring rigorous analytical precision.
- Spectral Library-Based DIA Quantification: Spectral library-based DIA workflows are becoming increasingly popular because of their excellent reproducibility and low inter-laboratory variability. Recent collaborative studies have reported coefficient of variation (CV) values below 15%, highlighting the robustness of this approach.
3. High-Resolution MS Platforms for Impurity Profiling of Biosimilars
Orbitrap and time-of-flight (ToF) systems are widely used in biosimilar analysis. Platform selection depends on resolution, scan speed, sensitivity, and workflow compatibility.
| Platform | Mass Accuracy | Resolution | Key Strength in Biosimilar Analysis |
|---|---|---|---|
| Orbitrap Astral | <1 ppm | >240,000 FWHM | Fast DIA analysis for HCPs and sequence variants |
| Q Exactive HF-X | <2 ppm | >120,000 FWHM | Industry standard for peptide mapping |
| Bruker timsTOF Pro 2 | <3 ppm | ~50,000 FWHM | Ion mobility separation for isobaric variants |
| Waters Xevo G3 QTof | <2 ppm | ~60,000 FWHM | Strong intact mass and glycan analysis |
| Thermo Orbitrap Eclipse | <2 ppm | >500,000 FWHM | Ultra-high-resolution intact antibody analysis |
Modern HRMS systems continue to improve in speed, sensitivity, and automation, supporting faster and more reliable biosimilar characterization.
Native Mass Spectrometry for Higher-Order Structure Analysis
Native MS preserves non-covalent interactions and higher-order protein structures. This approach allows scientists to:
- Resolve monomer and oligomer distributions
- Detect HCP co-purification
- Characterize half-antibody species
- Identify mispaired disulfide bonds
These capabilities are valuable because higher-order structural integrity directly affects biologic performance and immunogenicity.
Detailed analysis: Intact Mass Analysis for Biosimilars
4. Glycan Analysis in Impurity Profiling of Biosimilars
Glycosylation impurities such as high-mannose glycans, α-Gal, Neu5Gc, and truncated glycoforms are major regulatory concerns because they can influence immunogenicity and therapeutic performance.
The glycan profile of a biosimilar can change because of:
- Cell line selection
- Media composition
- Oxygen levels
- pH conditions
- Culture duration
For this reason, glycan comparability remains one of the most complex aspects of biosimilar development.
HRMS-Based Glycan Profiling Strategies
- Released N-Glycan Analysis: N-glycans are enzymatically released using PNGase F, fluorescently labeled using reagents such as 2-AB or RapiFluor-MS, and analyzed by HILIC-HRMS. This workflow supports simultaneous quantification of dozens of glycan species with excellent sensitivity and reproducibility.
- Glycopeptide Analysis: Tryptic glycopeptides are analyzed directly using LC-HRMS to obtain site-specific glycosylation information. This approach is particularly important for biologics containing multiple N-glycosylation sites because glycan occupancy and composition may differ significantly between domains.
- O-Glycan Profiling: O-glycan characterization typically requires specialized beta-elimination or enzymatic workflows such as OpeRATOR digestion. Advanced fragmentation techniques, including ETD and ECD, are often necessary for accurate site localization and structural assignment.
Comprehensive glycan analysis provides valuable insight into process consistency, product stability, and potential immunogenicity concerns. Because glycosylation is highly process-dependent, HRMS-based glycan profiling remains a central component of biosimilar analytical comparability packages.
Discover our Glycosylation Analysis of Biosimilars
5. Regulatory Expectations for Impurity Profiling of Biosimilars
Regulatory agencies expect HRMS impurity profiling data to support a comprehensive totality-of-evidence approach for biosimilarity assessment. Analytical similarity studies must demonstrate that the biosimilar and reference product are highly comparable across critical quality attributes using scientifically justified statistical and analytical methodologies.
The FDA’s 2019 analytical similarity guidance identifies several Tier 1 quality attributes that rely heavily on HRMS data. These attributes are evaluated using predefined equivalence criteria because of their direct impact on product quality and clinical performance.
Tier 1 Quality Attributes Supported by HRMS
- Charge variant profiling using CEX-MS
- Glycan profiling using HILIC-HRMS
- Peptide mapping and sequence verification
- Primary sequence confirmation
- Sequence variant analysis
Strong documentation, reproducibility, and orthogonal validation are essential for regulatory approval.
Common Regulatory Deficiencies
- Sequence coverage below 95%
- Lack of orthogonal confirmation for variants
- Missing site-specific glycosylation data
- Poor HCP method validation
- Inconsistent analytical sensitivity across lots
These deficiencies can result in additional information requests, regulatory delays, or concerns regarding biosimilar comparability. Therefore, analytical methods must be carefully optimized, validated, and documented according to current regulatory expectations.
Access our Regulatory Support for Generic Drug Development
6. Emerging Technologies Advancing Impurity Profiling of Biosimilars
Recent technological innovations in trapped ion mobility spectrometry (TIMS), real-time search (RTS), and artificial intelligence-assisted spectral analysis are transforming biosimilar impurity profiling workflows. These advancements are significantly reducing analysis time while improving sensitivity, selectivity, and confidence in impurity identification.
Modern HRMS platforms now combine multiple orthogonal separation dimensions with advanced computational algorithms, enabling more comprehensive characterization of highly complex biologic products. As biosimilars become increasingly sophisticated, these next-generation technologies are expected to play an even greater role in regulatory characterization strategies.
- 4D-Proteomics (PASEF + HRMS): Ion mobility introduces a fourth analytical dimension that improves separation of isobaric sequence variants and structurally similar species. This technology is particularly useful for resolving substitutions such as Ile/Leu variants or distinguishing deamidation events from isotope overlap artifacts.
- Real-Time Search (RTS) with Orbitrap: RTS enables on-the-fly database searching during data acquisition, allowing triggered MS3 fragmentation for co-fragmented variants. This approach improves sensitivity and analytical specificity without significantly increasing instrument runtime.
- Hydrogen-Deuterium Exchange MS (HDX-MS): HDX-MS detects subtle higher-order structural differences between biosimilars and reference products by monitoring hydrogen exchange kinetics. Regulatory agencies, particularly the EMA, increasingly view HDX-MS as a valuable orthogonal tool for evaluating conformational comparability and immunogenicity risk.
- AI-Assisted Variant Scoring: Machine learning algorithms trained on large biologic sequence datasets can assign probability scores to candidate sequence variants. These AI-driven approaches reduce false-positive identifications, improve data interpretation speed, and support high-throughput biosimilar screening programs.
The integration of artificial intelligence with HRMS bioinformatics is expected to substantially enhance analytical throughput and decision-making efficiency in future biosimilar characterization workflows.
Understand the Forced Degradation of Biosimilars
Conclusion: Advanced Impurity Profiling of Biosimilars Requires Scientific and Regulatory Expertise
The characterization of biosimilar impurities and sequence variants has become a critical requirement in modern biologic development. Today’s HRMS technologies can detect impurities at extremely low levels while supporting detailed analysis of glycosylation, sequence heterogeneity, and higher-order structures.
As regulatory expectations continue to evolve for monoclonal antibodies, fusion proteins, bispecific antibodies, and antibody-drug conjugates (ADCs), comprehensive Impurity Profiling of Biosimilars is becoming even more important. Success depends not only on advanced instrumentation but also on validated workflows, strong bioinformatics, and regulatory understanding.
The ongoing development of ion mobility, AI-assisted analysis, HDX-MS, and next-generation HRMS platforms is shaping the future of biosimilar characterization.
Explore our Charge Variant Analysis in Biosimilars
ResolveMass Laboratories Inc. combines expertise in HRMS platform science, biosimilar analytical strategy, and regulatory-focused bioinformatics to support global biosimilar development programs. Through scientifically rigorous workflows and submission-ready analytical studies, the company helps clients generate high-quality comparability packages for worldwide regulatory submissions.
📩 Ready to discuss your biosimilar characterization program?
Contact ResolveMass Laboratories Inc.
Frequently Asked Questions (FAQs)
ICH Q6B states that all product-related impurities and sequence variants in biosimilars must be properly identified, characterized, and evaluated for their possible effect on safety and efficacy. Although the guideline does not define a fixed numerical limit, regulatory agencies such as the FDA and EMA usually expect detailed analysis for variants detected around or above 0.1%. Advanced orthogonal mass spectrometry methods are commonly used to confirm these findings. Companies must also provide scientific justification for any detected variants and explain their potential clinical impact.
Data-independent acquisition (DIA) improves sequence variant analysis because it collects fragmentation data from all precursor ions within a selected mass range. In contrast, data-dependent acquisition (DDA) mainly focuses on the most abundant ions, which may cause low-level variants to be missed. DIA provides more consistent MS2 data across repeated injections, making it highly reliable for detecting variants present at very low levels. This approach improves reproducibility, sensitivity, and confidence in regulatory biosimilar studies.
High-resolution mass spectrometry (HRMS) is extremely valuable for identifying and quantifying specific host cell proteins, but it does not completely replace ELISA in regulatory submissions. ELISA is still widely accepted as the standard method for total HCP testing during GMP release. However, HRMS offers important advantages because it can detect individual HCP contaminants that may not be visible through ELISA methods. Most regulatory agencies now recommend using both ELISA and LC-MS/MS together to achieve a more complete impurity assessment.
Regulatory agencies generally expect peptide mapping studies to achieve at least 95% sequence coverage for biosimilars. However, many advanced laboratories aim for sequence coverage above 99% by using multiple enzyme digestion strategies such as Trypsin, GluC, and Lys-C. Regions with poor coverage can create regulatory concerns, especially in critical antibody domains or glycosylation sites. If any sections remain uncovered, additional analytical methods are usually required to confirm the complete protein sequence.
Isoleucine and leucine have identical molecular weights, which makes them difficult to distinguish using standard mass spectrometry alone. Specialized fragmentation methods such as ETD and ECD are commonly used because they generate fragment ions that help differentiate these amino acids. Ion mobility spectrometry can also assist by separating peptides based on structural differences. These advanced analytical approaches improve confidence in sequence variant identification during biosimilar characterization studies.
Native mass spectrometry analyzes proteins under non-denaturing conditions, allowing scientists to study intact protein structures and non-covalent interactions. This method helps evaluate quaternary structure, aggregation behavior, and ligand binding properties in biosimilars. Native MS is especially useful for monoclonal antibodies, Fc-fusion proteins, and bispecific antibodies where structural integrity is critical. Regulatory reviewers increasingly consider native MS as supportive evidence for higher-order structural comparability.
Glycosylation impurities such as Neu5Gc and α-Gal are commonly analyzed using released glycan profiling or glycopeptide mapping with HILIC-HRMS. These impurities can be accurately detected because they show specific mass differences compared to normal human glycans. Modern Orbitrap-based HRMS platforms can identify these glycan variants at extremely low concentrations. Their measurement is important because non-human glycans may increase immunogenicity risk in patients receiving biologic therapies.
Several advanced software platforms are used for biosimilar sequence variant analysis and HRMS data interpretation. Commonly used tools include Byonic, Proteome Discoverer, Mascot, and BioPharma Finder. For DIA-based workflows, platforms such as Spectronaut and DIA-NN are widely used because of their strong quantitative performance. Regulatory submissions often include validation data for these software workflows to demonstrate accuracy, sensitivity, and controlled false discovery rates.
Reference:
- Segu, Z., Stone, T., Berdugo, C., Roberts, A., Doud, E., & Li, Y. (2020). A rapid method for relative quantification of N-glycans from a therapeutic monoclonal antibody during trastuzumab biosimilar development. MAbs, 12(1), 1750794. https://pmc.ncbi.nlm.nih.gov/articles/PMC7188402/
- Nupur, N., Joshi, S., Guillarme, D., & Rathore, A. S. (2022). Analytical similarity assessment of biosimilars: Global regulatory landscape, recent studies and major advancements in orthogonal platforms. Frontiers in Bioengineering and Biotechnology, 10, 832059. https://pmc.ncbi.nlm.nih.gov/articles/PMC8865741/
- D’Atri, V., Guillarme, D., & Beck, A. (2025). Biopharmaceutical analysis—current analytical challenges, limitations, and perspectives. Analytical and Bioanalytical Chemistry. https://pmc.ncbi.nlm.nih.gov/articles/PMC12783247/
- Berkowitz, S. A., Engen, J. R., Mazzeo, J. R., & Jones, G. B. (2012). Analytical tools for characterizing biopharmaceuticals and the implications for biosimilars. Nature Reviews Drug Discovery, 11(7), 527–540. https://pmc.ncbi.nlm.nih.gov/articles/PMC3714370/



