Introduction:
Native Mass Spectrometry for Biosimilars has emerged as a highly advanced analytical strategy for investigating the quaternary structure and non-covalent assemblies of therapeutic proteins that are often invisible under conventional denaturing conditions. By transferring biomolecules from a solution that closely mimics physiological conditions into the gas phase while preserving their folded conformations, this technology allows accurate assessment of stoichiometry, binding interactions, and higher-order structural integrity. These characteristics are critical when demonstrating analytical similarity between a biosimilar candidate and its reference biologic.
Share via:
Article Summary:
- Native Mass Spectrometry (MS) for biosimilars enables analysis of therapeutic proteins in their folded, near-physiological state while preserving non-covalent interactions and quaternary structures.
- Unlike denaturing MS, native MS focuses on higher-order structure (HOS), intact molecular assemblies, stoichiometry, and binding interactions that directly impact biologic function and efficacy.
- Native MS provides accurate measurement of intact protein complexes, helping confirm biosimilar similarity in terms of assembly state, subunit composition, conformational integrity, and ligand binding.
- The technique generates lower charge states and higher m/z values compared with denaturing MS, resulting in clearer spectra with reduced overlap between glycoforms and protein variants.
- Native MS plays a critical role in biosimilar characterization by identifying subtle structural heterogeneity, low-abundance variants, chain mispairing in bispecific antibodies, and drug-to-antibody ratio (DAR) distributions in ADCs.
- Regulatory agencies such as the FDA and EMA increasingly recognize native MS as an important component of the “totality of evidence” approach for demonstrating biosimilarity and reducing residual uncertainty.
- Successful native MS analysis requires optimized instrumentation, high-mass-range configurations, volatile buffer systems like ammonium acetate, and advanced deconvolution workflows for accurate intact mass interpretation.
- Native MS is often combined with complementary techniques such as SEC-MALS, AUC, HDX-MS, and ion mobility MS to deliver comprehensive structural characterization, impurity profiling, and advanced biosimilar comparability assessment.
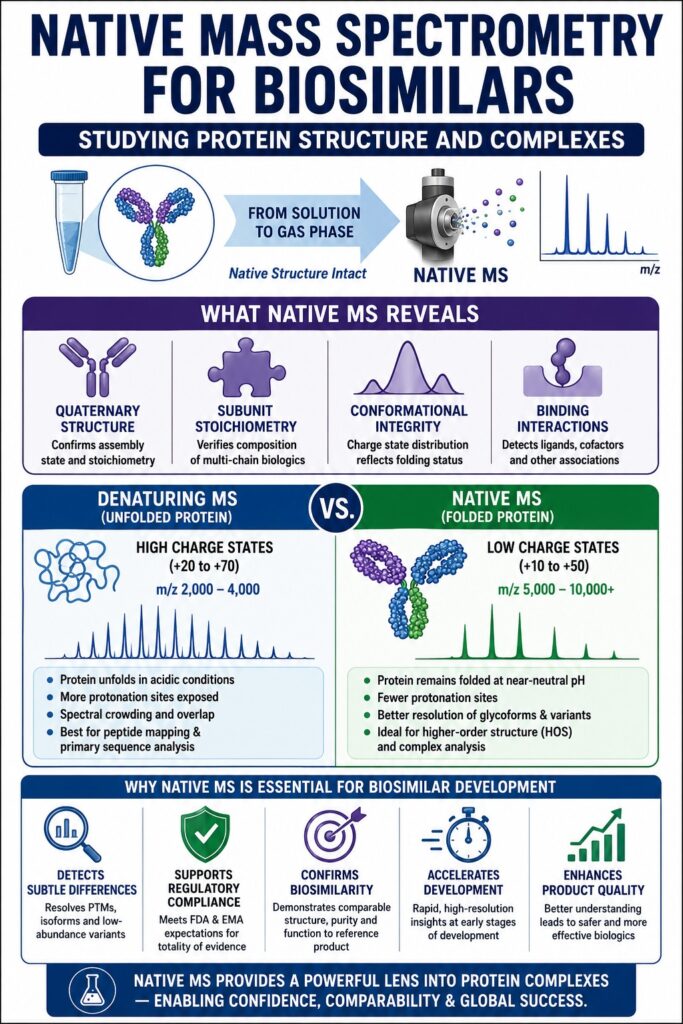
Advanced Structural Characterization Through Native Mass Spectrometry for Biosimilars
Native Mass Spectrometry for Biosimilars plays a central role in evaluating the functional and folded state of therapeutic proteins by maintaining non-covalent interactions throughout the liquid-to-gas phase transition. Unlike denaturing MS approaches that primarily focus on amino acid sequence determination, native MS emphasizes higher-order structural (HOS) organization and delivers precise intact mass measurements for complete molecular assemblies that directly influence biological activity and therapeutic efficacy.
Learn more about the full scope of analysis: Explore Comprehensive Biosimilar Characterization Using Mass Spectrometry
Preservation of Quaternary Structure
One of the key outputs generated through native MS is the accurate mass measurement of the intact protein complex. This information reveals the assembly state and stoichiometry of the molecule. In biosimilar development, this is especially important for confirming that the quaternary structure, meaning the arrangement of multiple folded subunits, matches that of the reference product.
For instance, a therapeutic monoclonal antibody (mAb) may predominantly exist as a 150 kDa monomer while also containing trace quantities of non-covalent dimers. Native MS can quantify these species with a level of precision that exceeds many chromatographic approaches such as SEC-UV.
This preservation of structural integrity is achieved by using volatile aqueous buffers, commonly ammonium acetate, under near-neutral pH conditions. During electrospray ionization (ESI), the protein remains folded, resulting in a smaller solvent-exposed surface area (SASA). As a consequence, fewer protonation sites are available, producing ions with lower charge states and higher $m/z$ values.
The resulting charge state distribution (CSD) becomes a highly sensitive indicator of conformational integrity. A narrow CSD generally reflects a compact and properly folded protein, whereas a broader distribution may indicate structural heterogeneity or partial unfolding.
Deep dive into intact molecular analysis: View Detailed Intact Mass Analysis for Biosimilars
Direct Measurement of Subunit Stoichiometry
Native Mass Spectrometry for Biosimilars enables precise determination of the number and ratio of unique subunits present within a protein complex. This capability is essential for confirming that a biosimilar reproduces the same molecular composition as the originator product.
By comparing the intact holo-complex mass measured under native conditions with the masses of individual subunits obtained under denaturing conditions, scientists can accurately determine subunit composition. This becomes particularly valuable for fusion proteins and multi-component biologics such as cytokine-receptor traps or hexameric IgG structures.
Native MS also distinguishes true binding interactions from simple co-purification artifacts. This distinction is critical during process development in laboratories such as ResolveMass Laboratories Inc., where confirming assembly purity is necessary for regulatory compliance. The ability to characterize associated proteins, nucleic acids, glycans, or lipids provides a more complete understanding of the structural biology of the therapeutic molecule.
| Feature | Impact on Biosimilarity Assessment | Mechanism of Detection |
|---|---|---|
| Holo-complex Mass | Confirms global assembly state | High $m/z$ Orbitrap acquisition |
| Subunit Stoichiometry | Verifies composition of multi-chain biologics | Comparison of native and denatured masses |
| Charge State Distribution | Evaluates conformational integrity and folding | Peak width and charge distribution analysis |
| Ligand Binding | Measures cofactor or small-molecule occupancy | Mass shifts observed in native spectra |
Comparative Analysis: Native MS vs. Denaturing MS
Native Mass Spectrometry for Biosimilars provides a significantly clearer spectral profile for heterogeneous biomolecules because reduced charge states minimize overlap between glycoforms and proteoforms in the m/z domain. While denaturing MS remains the standard for peptide mapping and primary sequence confirmation, native MS uniquely resolves intact antibody complexity without interference from overlapping charge envelopes.
Compare analytical methodologies: Prove Biosimilarity Using LC-MS Workflows
Charge State and m/z Shifts in Monoclonal Antibodies
Under denaturing ESI-MS conditions, monoclonal antibodies are exposed to acidic environments, commonly 0.1% formic acid, along with organic solvents such as acetonitrile. These conditions unfold the protein, exposing basic residues like lysine, arginine, and histidine to protonation.
As a result, high charge states are generated, typically ranging from +20 to +70 for a 150 kDa antibody. These ions appear in the lower $m/z$ range of approximately 2,000 to 4,000. Although beneficial for sensitive detection, this often causes substantial spectral crowding, where glycan variants from neighboring charge states overlap.
Native MS changes this analytical landscape completely. By preserving proteins in their folded, neutral-pH state, the number of protonation events decreases, resulting in charge states between +10 and +50. Consequently, ions shift into the higher $m/z$ region of 5,000 to 10,000+.
This separation greatly improves spectral spatial resolution. Glycoforms such as G0F/G0F, G0F/G1F, and G1F/G1F become more distinguishable at the intact molecular level, enabling more reliable quantification without extensive computational deconvolution.
Resolving Heterogeneity: Infliximab Case Study
Studies involving the biosimilar infliximab demonstrate the practical value of Native Mass Spectrometry for Biosimilars when comparing originators and biosimilar candidates. Analyses involving commercial originators, marketed biosimilars, and in-house CHO-derived candidates revealed that native MS produced cleaner profiles of C-terminal lysine heterogeneity.
Infliximab commonly contains three major variants defined by heavy-chain C-terminal lysines: 0K, 1K, and 2K.
Under native conditions, the 1xG0F_G0F glycoform was measured at approximately 27.37% abundance in the originator product, showing strong agreement with denaturing MS data. However, native MS offered superior visualization of low-abundance variants, including +2 lysine species that may otherwise be hidden by spectral overlap or background noise under denaturing conditions.
This enhanced clarity is vital for scientists at ResolveMass Laboratories Inc. when confirming that the biosimilar fingerprint falls within the acceptable variability range of the reference product.
Analyze complex charge distributions: Read More on Charge Variant Analysis in Biosimilars
| MS Condition | $m/z$ Range (mAb) | Charge States | Primary Application | PTM Resolution |
|---|---|---|---|---|
| Denaturing | 2,000 – 4,000 | +20 to +70 | Primary sequence and glycan profiling | High with ultra-HRAM |
| Native | 5,000 – 10,000+ | +10 to +50 | HOS, quaternary structure, aggregate analysis | Superior due to reduced overlap |
Regulatory Expectations and the Role of Native Mass Spectrometry for Biosimilars
Native Mass Spectrometry for Biosimilars is increasingly recognized by regulatory authorities such as the U.S. Food and Drug Administration and the European Medicines Agency as an essential component of the “totality of evidence” approach used to establish biosimilarity.
Because biologics naturally exhibit variability in glycosylation, folding, and post-translational modifications, exact duplication of the originator is impossible. Regulators therefore require comprehensive multidimensional characterization to demonstrate that no clinically meaningful differences exist.
ICH Q5E and the Totality of Evidence
The ICH Q5E guideline establishes the scientific framework for evaluating how manufacturing changes or independent production processes affect biologic quality, safety, and efficacy.
Biosimilar developers must prove that the molecular profile of the biosimilar remains within the statistical variability range established from multiple lots of the reference product. Native MS contributes a highly informative “fingerprint-like” assessment that simultaneously evaluates several critical quality attributes (CQAs), including:
- Primary sequence integrity
- Higher-order structure
- Purity characterization
- Complex assembly integrity
The totality-of-evidence model means that strong analytical similarity can reduce the requirement for extensive clinical efficacy trials. When native MS demonstrates that a biosimilar closely matches the reference product in its intact native state, regulatory uncertainty surrounding clinical safety and efficacy is substantially reduced.
Identify and monitor essential attributes: Understand Critical Quality Attributes (CQAs) in Biosimilars
FDA and EMA Characterization Perspectives
Although both agencies require products to be “highly similar,” their emphasis differs slightly.
The FDA focuses on the absence of clinically meaningful differences in safety, potency, and purity. The EMA places greater emphasis on building a complete analytical narrative covering structure, biological activity, and efficacy.
Native Mass Spectrometry for Biosimilars is uniquely positioned to satisfy both perspectives. The technology can resolve oligomeric distributions, identify trace host-cell protein co-purification, and characterize intact molecular assemblies in detail.
At ResolveMass Laboratories Inc., specialized HRMS workflows are designed specifically to support IND, BLA, and MAA submissions in alignment with global regulatory requirements.
Compare biosimilar lots for regulatory approval: Learn About Biosimilar Comparability Studies
| Regulatory Concept | Definition / Requirement | Native MS Contribution |
|---|---|---|
| Highly Similar | Minor differences permitted in inactive components | Confirms matching folding and HOS |
| Totality of Evidence | Combined analytical, functional, and clinical data | Forms analytical foundation |
| Residual Uncertainty | Unresolved differences after testing | Resolves quaternary structure concerns |
| Analytical Fingerprinting | Simultaneous assessment of multiple CQAs | Maps PTMs on intact complexes |
Characterizing Multispecific Antibodies and Complexes
Native Mass Spectrometry for Biosimilars is indispensable for evaluating multispecific antibodies (MsAbs) and antibody-drug conjugates (ADCs). The technique can quantify chain mispairing and payload distribution while preserving the non-covalent or covalent interactions that define these highly complex therapeutic modalities.
Resolving Chain Mispairing in Bispecific Antibodies
The co-expression of multiple heavy and light chains during multispecific antibody production often generates unwanted mispaired species, including homodimers, half-antibodies, and incorrectly assembled trispecifics.
Because these variants can possess nearly identical masses to the intended product, native MS becomes essential for differentiating them based on intact complex mass and subunit composition.
Native MS also supports early-stage CHO clone screening, enabling researchers to identify clones with lower mispairing frequencies before scale-up manufacturing begins.
At ResolveMass Laboratories Inc., SEC-native MS workflows are used to analyze clarified samples directly, eliminating the need for time-consuming protein A purification during early development.
DAR Analysis in ADCs
For ADCs, the drug-to-antibody ratio (DAR) is one of the most important quality attributes influencing therapeutic efficacy and toxicity.
Native MS accurately resolves the mass changes associated with individual payload additions while preserving the antibody assembly. Native top-down mass spectrometry (nTD-MS) further enables fragmentation of the intact ADC within the instrument itself, allowing researchers to determine both average DAR and precise conjugation sites.
Combining native ionization with proton-transfer charge reduction (PTCR) further decreases spectral congestion and enhances the detection of high-mass fragments.
| Modality | Structural Challenge | Native MS Advantage |
|---|---|---|
| Bispecific Antibodies | Chain mispairing and homodimers | Intact mass and stoichiometry analysis |
| ADCs | DAR distribution and payload stability | High-resolution intact mass profiling |
| Fusion Proteins | Misfolding and truncation | Charge state distribution monitoring |
| Immune Complexes | Antigen-antibody binding ratios | Non-covalent stoichiometry analysis |
Technical Parameters and Optimization
Successful implementation of Native Mass Spectrometry for Biosimilars requires specialized instrumentation and precise optimization to ensure stable transmission and desolvation of high-mass ions while preserving native structural features.
Mass Range and Pressure Optimization
Because folded proteins acquire fewer charges, native ions appear at much higher $m/z$ values than denatured proteins. Instruments therefore require extended high-mass-range configurations.
Key instrument requirements include:
- Low-frequency quadrupoles capable of transmitting ions beyond $m/z$ 8,000–30,000
- Elevated buffer gas pressure for collisional cooling
- Controlled in-source dissociation (SID) for solvent removal without disrupting complexes
Deconvolution and Data Processing
Native MS spectra often contain broad peaks and unresolved isotopic patterns due to residual solvent and adduct formation. As a result, deconvolution software such as ReSpect™ is preferred over isotopically focused algorithms like Xtract™.
At ResolveMass Laboratories Inc., BioPharma Finder™ 3.0 is used to process native datasets with optimized parameters for intact mAbs, typically within a 140,000–160,000 Da output range.
| Tune Parameter | Denaturing Setting | Native Setting | Purpose |
|---|---|---|---|
| S-lens RF Voltage | 80 | 200 | Improves high-mass ion transmission |
| Capillary Temperature | 275 °C | 275 °C | Maintains desolvation without unfolding |
| Probe Heater Temperature | 175 °C | 275 °C | Accelerates evaporation of aqueous buffers |
| SID Voltage | 100 | 150 | Removes solvent and buffer adducts |
| MS Mode | Protein Mode | HMR Mode | Extends measurable $m/z$ range |
Sample Preparation and Buffer Exchange
Native Mass Spectrometry for Biosimilars depends heavily on effective sample preparation, particularly the removal of non-volatile salts such as sodium chloride and their replacement with volatile buffers that preserve protein integrity.
Importance of Ammonium Acetate
Ammonium acetate is considered the preferred native MS buffer because it is volatile and preserves native protein conformation while minimizing adduct formation.
Typical concentrations range between 50 mM and 200 mM under near-neutral pH conditions.
In some cases, charge-reducing additives such as triethylammonium acetate (TEAA) or imidazole are introduced to stabilize complexes or further improve spectral resolution.
Online and Offline Buffer Exchange Strategies
Offline Exchange
Traditional spin-column approaches require approximately 25–30 minutes per sample. Although effective, these methods are labor-intensive and may lead to dilution or sample loss.
Online Buffer Exchange (OBE)
Automated systems such as DynaChip X1™ and online SEC-MS perform buffer exchange seconds before ionization. These methods are significantly faster and frequently produce cleaner spectra because proteins spend less time in volatile buffer environments before analysis.
ResolveMass Laboratories Inc. utilizes both workflows depending on the complexity and stability of the biosimilar candidate.
| Preparation Method | Time Per Sample | Advantages | Limitations |
|---|---|---|---|
| Manual Spin Filter | 30–60 min | Cost-effective and simple | Sample loss and aggregation risk |
| Offline Dialysis | 4–24 hours | Gentle on complexes | Slow and high-volume |
| nSEC-MS | 15–28 min | Automated and separates aggregates | Potential sample dilution |
| Rapid OBE | <5 min | Extremely fast and integrated | Requires specialized microfluidics |
Impurity Profiling and Higher-Order Structure Analysis
Native Mass Spectrometry for Biosimilars provides detailed characterization of both product-related and process-related impurities that may affect therapeutic safety and efficacy.
Detecting Product-Related Variants
Product-related impurities include:
- Truncated variants
- Glycation products
- Oxidized species
- Non-covalent aggregates
Native MS excels at resolving high-molecular-weight species such as dimers and trimers while preserving their assembly state.
The technology also provides global insight into disulfide integrity. Mispaired disulfide bonds may alter charge state distributions or generate detectable intact mass shifts compared with the reference product.
Identify contaminants in your samples: Read About Impurity Profiling of Biosimilars
Monitoring Process-Related Impurities
Host-cell proteins (HCPs), leachables, and extractables are additional regulatory concerns.
Although LC-MS/MS with DIA remains the primary technique for low-level HCP analysis, native MS offers rapid at-line detection of co-purified contaminants associated with the therapeutic complex.
At ResolveMass Laboratories Inc., advanced HRMS workflows are specifically designed to identify manufacturing-derived contaminants such as silicone oils, tubing residues, and filter leachables.
| Impurity Category | Example | Native MS Capability |
|---|---|---|
| Product-Related | Non-covalent aggregates | Resolves oligomeric species |
| Product-Related | Truncated molecules | Detects intact mass differences |
| Product-Related | Mispaired disulfides | Alters CSD and intact masses |
| Process-Related | Co-purified HCPs | Detects non-target complexes |
| Process-Related | Leachables and adducts | Identifies mass additions |
Orthogonal Comparison: Native MS vs. SEC-MALS and AUC
Native Mass Spectrometry for Biosimilars achieves maximum effectiveness when combined with orthogonal analytical approaches.
SEC-MALS
SEC-MALS measures absolute molecular mass and aggregate content independent of column calibration.
Limitation
SEC-MALS cannot resolve subtle variants such as individual glycoforms or +1 lysine differences.
Complementary Strength
Native MS adds precise mass identity and structural detail to SEC-MALS findings.
AUC and Mass Photometry
Analytical ultracentrifugation (AUC) remains a gold standard for aggregate analysis because it is matrix-free and operates directly in formulation buffer.
Mass photometry provides exceptionally rapid analysis with low sample consumption.
At ResolveMass Laboratories Inc., native MS is integrated alongside HDX-MS, SEC-MALS, and AUC to support advanced biosimilar fingerprinting strategies.
| Technique | Principle | Primary Advantage |
|---|---|---|
| Native MS | Gas-phase mass analysis | High-resolution PTM and complex characterization |
| SEC-MALS | Light scattering | Absolute mass in formulation buffer |
| AUC | Sedimentation velocity | Matrix-free aggregate quantification |
| Mass Photometry | Single-molecule scattering | Extremely rapid screening |
| HDX-MS | Deuterium exchange | Maps folding and interaction surfaces |
Case Studies: High-Resolution Biosimilar Profiling
Native Mass Spectrometry for Biosimilars has demonstrated exceptional performance in originator-versus-biosimilar comparisons involving major biologics such as trastuzumab, cetuximab, and infliximab.
Trastuzumab Comparisons
Native MS successfully resolved multiple trastuzumab glycoforms including:
- G0/G0F
- G0F/G0F
- G0F/G1F
- G1F/G1F
- G1F/G2F
Mass accuracy remained within ±0.8–1.1 Da for a 148 kDa molecule.
Ion mobility MS further identified low-abundance dimeric species in certain biosimilar lots that were absent in the originator.
Cetuximab and Infliximab Studies
Cetuximab presents extensive Fab-region glycosylation, increasing heterogeneity substantially. Even so, native MS remains one of the most effective approaches for intact molecular profiling.
For infliximab, advanced 2D-LC coupled with native MS enabled detection of 23 chemically unique fragments in a single sample, demonstrating the remarkable resolving capability of modern HRMS systems.
Future Outlook: Native Top-Down MS and Automation
The future of Native Mass Spectrometry for Biosimilars is closely tied to advances in fragmentation technologies and high-throughput automation.
Native Top-Down Fragmentation
Native top-down MS enables direct sequencing of folded proteins using approaches such as:
- 213 nm UVPD
- Electron Transfer Dissociation (ETD)
These methods often provide superior sequence coverage for disulfide-rich regions compared with denaturing workflows.
Automated High-Throughput Platforms
Chip-based nano-ESI systems such as NanoMate have dramatically improved analysis speed and reproducibility.
These platforms support:
- Fragment-based drug discovery (FBDD)
- Large-scale biosimilarity screening
- Rapid batch-to-batch comparisons
At ResolveMass Laboratories Inc., automated Orbitrap-based workflows combined with high-throughput sample preparation are being used to deliver next-generation structural characterization for complex biotherapeutics.
Conclusion: Achieving Global Regulatory Compliance Through Advanced Structural MS
Native Mass Spectrometry for Biosimilars represents one of the most advanced structural characterization technologies available in the modern biopharmaceutical industry. By resolving quaternary assemblies, higher-order structural variants, and subtle conformational differences that define biological identity, native MS enables manufacturers to satisfy the rigorous “highly similar” requirements established by the FDA and EMA.
As therapeutic biologics continue to increase in structural complexity, high-resolution native-state mass spectrometry will remain essential for biosimilar development, analytical comparability, and regulatory success.
ResolveMass Laboratories Inc. continues to lead this field by combining technical expertise, advanced HRMS workflows, and regulatory insight to transform highly complex molecular data into clinically meaningful conclusions.
For more information regarding advanced structural characterization services and biosimilar development support, please contact the scientific team at:
FAQs on Native Mass Spectrometry for Biosimilars
Native mass spectrometry maintains the natural folded conformation of proteins along with their non-covalent interactions during analysis. This makes it possible to study higher-order structures, molecular assembly, and protein complex formation that are typically disrupted under denaturing conditions. As a result, researchers gain a more accurate understanding of the biologic’s functional structure.
Ammonium acetate is the most frequently used volatile buffer in native MS workflows because it preserves protein stability while evaporating cleanly during ionization. It is generally used at concentrations between 50 mM and 200 mM under near-neutral pH conditions. This buffer minimizes salt adduct formation and helps maintain native protein conformation throughout analysis.
Regulatory agencies favor fingerprint-style characterization because it evaluates multiple product attributes simultaneously using complementary analytical methods. This multidimensional assessment reduces uncertainty regarding biosimilarity and improves confidence that the biosimilar closely matches the reference biologic. Strong analytical evidence may also reduce the need for extensive clinical testing.
High Mass Range (HMR) mode expands the detectable m/z range of the mass spectrometer, allowing it to analyze large native protein complexes with low charge states. Without HMR capability, many intact biologics and non-covalent assemblies would fall outside the instrument’s measurable range. This feature is especially important for studying monoclonal antibodies and high-molecular-weight complexes.
Before native MS analysis, samples undergo buffer exchange to replace non-volatile salts with volatile buffers such as ammonium acetate. This process may be performed using spin columns, dialysis methods, or automated online systems including SEC-MS and DynaChip X1™ platforms. Proper salt removal is essential for improving spectral quality and mass accuracy.
Post-translational modifications such as glycosylation create small mass differences that can influence protein structure and biological activity. Native MS improves the resolution of these PTMs because lower charge states reduce spectral overlap between glycoforms and proteoforms. This enables more accurate profiling and quantification of intact molecular variants.
Reference:
- National Center for Biotechnology Information. (2024). Reverse engineering and characterization of long-acting injectable microspheres: Current approaches and challenges. PubMed Central. https://pmc.ncbi.nlm.nih.gov/articles/PMC10892806/
- European Medicines Agency. (2025). Reflection paper on a tailored clinical approach in biosimilar development (EMA/CHMP/BMWP/60916/2025). https://www.ema.europa.eu/en/documents/other/reflection-paper-tailored-clinical-approach-biosimilar-development_en.pdf
- National Center for Biotechnology Information. (2023). Recent advances in long-acting injectable microsphere formulations: Design, characterization, and challenges. PubMed Central. https://pmc.ncbi.nlm.nih.gov/articles/PMC10432323/